MBD4 (methyl-CpG binding domain protein 4)
2011-07-01 Huan X Meng , Richard R Meehan AffiliationMRC Human Genetics Unit, Western General Hospital, Crewe Road, Edinburgh EH4 2XU, UK
DNA/RNA
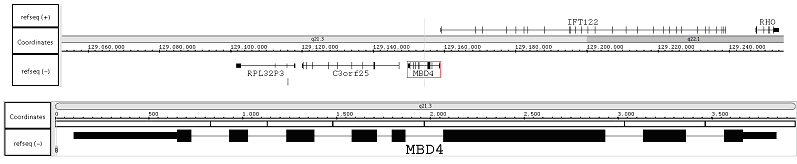
Description
MBD4 (NM_003925) is a gene of 9060 bp coded by 8 exons from 129158852 to 129149792 according to GRCh37 human genome assembly v37 by IGB browser.
Transcription
The transcript encoded by NM_003925.1, mRNA length of 2470 bp.
Proteins
Note
MBD4 has two significant domains identified by homology and functional analysis: a highly conserved glycosylase domain (60% conservation between species) at the C-terminus, and a less well conserved methyl-CpG binding (MBD) domain (37% conservation between species) at the N-terminus (Bellacosa et al., 1999; Hendrich and Bird, 1998; Meng et al., 2011). The domain structure of MBD4 indicates that the gene could have been formed from the fusion of a glycosylase motif with a methyl-CpG binding domain (Hendrich and Tweedie, 2003). The N-terminal MBD domain of MBD4 is able to discriminate between methylated and unmethylated double stranded DNA (Hendrich and Bird, 1998). The N-terminal glycosylase domain of MBD4 was shown in vitro to recognise and repair the mismatched products from deamination including Thymine and Uracil (Hendrich et al., 1999; Wong et al., 2002), thus it has been proposed to take part in active demethylation. The methyl-CpG binding domain of MBD4 binds preferentially to m5CpG x TpG mismatches - the primary product of deamination at methyl-CpG.
It has been proposed that MBD4 proteins arose as a fusion protein between MBD and glycosylase domain ancestors in the vertebrate lineage, and MBD2/3 represents the ancestral methyl-CpG binding protein (Hendrich and Tweedie, 2003). Interestingly, a prototypical MBD4 protein and its putative ancestor MBD2/3 were identified outside vertebrates in the cephalochordate amphioxus (Branchiostoma floridae) (Albalat, 2008), in addition a putative MBD4 gene was predicted in the Ciona intestinalis genome, but this lacks an MBD domain. The finding of a putative MBD4 protein in this invertebrate-vertebrate transition model organism pushes back the origin of MBD4 proteins in evolutionary time, and argues against the fusion hypothesis of origin of MBD4 proteins (Albalat, 2008). The MBD domain of MBD4 is most similar to that of MeCP2 in primary sequence (Ballestar and Wolffe, 2001; Hendrich and Bird, 1998).
It has been proposed that MBD4 proteins arose as a fusion protein between MBD and glycosylase domain ancestors in the vertebrate lineage, and MBD2/3 represents the ancestral methyl-CpG binding protein (Hendrich and Tweedie, 2003). Interestingly, a prototypical MBD4 protein and its putative ancestor MBD2/3 were identified outside vertebrates in the cephalochordate amphioxus (Branchiostoma floridae) (Albalat, 2008), in addition a putative MBD4 gene was predicted in the Ciona intestinalis genome, but this lacks an MBD domain. The finding of a putative MBD4 protein in this invertebrate-vertebrate transition model organism pushes back the origin of MBD4 proteins in evolutionary time, and argues against the fusion hypothesis of origin of MBD4 proteins (Albalat, 2008). The MBD domain of MBD4 is most similar to that of MeCP2 in primary sequence (Ballestar and Wolffe, 2001; Hendrich and Bird, 1998).
Description
The longest Human MBD4 protein has 580 amino acids, at approximately 66 kDa.
Expression
MBD4 expressed in all tissues tested with highest levels in Tonsil and Early Erythroid (CD71+) (BioGPS).
Localisation
MBD4 is a nuclear protein. Overexpressed MBD4-GFP co-localises to heterochromatin sites in mouse cells and can recruit partner proteins e.g. MLH1 (Hendrich and Bird, 1998; Ruzov et al., 2009).
Function
MBD4 is a methyl-CpG binding glycosylase (Bellacosa et al., 1999) that can excise and repair both C-T and C-U mutations at methylated and non-methylated CpGs via its glycosylase domain and adjacent binding site (Hendrich et al., 1999). Novel interacting partners of MBD4 include the mis-match repair (MMR) protein MLH1 (Bellacosa et al., 1999) and Fas-associated death domain (FADD) protein (Screaton et al., 2003), suggesting a potential link between genome surveillance and apoptosis. Consistent with these observations, reduced apoptosis occurs in the small intestine of mbd4-/- mice in response to a variety of DNA-damaging agents (Cortellino et al., 2003), and increased gastrointestinal tumorigenicity was observed for mbd4-/- mice on a tumor-susceptible Apc min background (Millar et al., 2002). Although Mbd4 inactivation did not increase mini-satellite instability (MSI) in the mouse genome, it did result in a 2- to 3-fold increase in C-to-T transition mutations at CpG sequences in splenocytes and epithelial cells of the small intestinal mucosa. The change in the gastrointestinal cancer phenotype was associated with an increase in somatic C-to-T mutations at CpG sites within the coding region of the wild-type Apc allele. These studies indicate that Mbd4 inactivation may alter the mutation spectrum in cancer cells and modify the cancer predisposition phenotype.
MBD4 can repress several methylated reporter genes (Fukushige et al., 2006; Kondo et al., 2005). However the global genomic targets of MBD4 have not been characterised. MBD4 knock out mice are viable and fertile, and show only mild physiological defects (Millar et al., 2002). However, morpholino knock-down of xMbd4 in Xenopus laevis shows serious developmental defects and fails to develop to a tadpole. Re-introduction of mouse Mbd4 mRNA can rescue the phenotype (Ruzov et al., 2009). There is increased C-T mutation rate observed in the Mbd4 knock out mice as a consequence of reduced glycosylase activity (Millar et al., 2002).
MBD4 can interact directly with both DNMT1 and MLH1 leading to recruitment of all three components at sites of DNA damage (Ruzov et al., 2009). The co-localization of DNMT1, MBD4 and MLH1 suggests that they may participate in a cellular checkpoint that monitors potential DNA hypomethylation events by detecting the presence or absence of the maintenance methyltransferase Dnmt1, perhaps at or adjacent to the replication fork. The recruitment of these components in response to localized DNA damage suggests that can have a role in the cellular decision whether to repair the lesion or activate apoptosis (Ruzov et al., 2009). Decreases in DNMT1 protein levels in ES cells and normal differentiated human cells can result in mismatch repair (MMR) defects (Guo et al., 2004; Kim et al., 2004; Loughery et al., 2011; Wang and James Shen, 2004) In the hTERT-1604 fibroblast cell line, a normal diploid cell line immortalised by telomerase over-expression, depletion of DNMT1 is sufficient to cause MMR defects and increased mutation rates at a CA17 microsatellite. This is associated with decreases in MMR protein levels, including MBD4: following activation of the DNA damage response (DDR). Blocking the DDR, and in particular PARP over-activation, also increases survival of the DNMT1 knockdowns (Loughery et al., 2011).
zMbd4 has been proposed to be a candidate protein that is involved in DNA demethylation activity in Zebrafish (Rai et al., 2008). Rai and colleagues showed MBD4 removes G:T mismatch-specific thymines, resulting from deamination of 5-methylcytosine (5-meC) via cytidine deaminase family members (Activation Induced deaminase (AID) and Apolipo-protein B RNA-editing catalytic component (Apobec)). Interestingly, deamination activity by AID/Apobec did not occur unless MBD4 and/or other possible factor are present and/or activated, and a catalytically inactive hMBD4 derivative (D560A) stabilized the putative G:T intermediate and prevented rapid thymine removal (Rai et al., 2008). MBD4 was also reported to interact with Xenopus DNMT1 to promote p53 dependent apoptosis (Ruzov et al., 2009), and DNMT3b, which was is proposed to methylate cytosine and to deaminate 5-meC, relying on an inefficient deaminase activity (Kangaspeska et al., 2008; Metivier et al., 2008). Recent studies in mice have implicated thymine DNA-glycosylase (TDG) in active DNA demethylation, TDG interacts with the deaminase AID and the damage response protein GADD45a. These findings suggest a two-step mechanism for DNA demethylation in mammals, whereby 5-methylcytosine and 5-hydroxymethylcytosine are first deaminated by AID to thymine and 5-hydroxymethyluracil, respectively, followed by TDG-mediated thymine and 5-hydroxymethyluracil excision repair (Cortazar et al., 2011; Cortellino et al., 2011). It is possible that MBD4 may make a contribution to this process.
MBD4 can repress several methylated reporter genes (Fukushige et al., 2006; Kondo et al., 2005). However the global genomic targets of MBD4 have not been characterised. MBD4 knock out mice are viable and fertile, and show only mild physiological defects (Millar et al., 2002). However, morpholino knock-down of xMbd4 in Xenopus laevis shows serious developmental defects and fails to develop to a tadpole. Re-introduction of mouse Mbd4 mRNA can rescue the phenotype (Ruzov et al., 2009). There is increased C-T mutation rate observed in the Mbd4 knock out mice as a consequence of reduced glycosylase activity (Millar et al., 2002).
MBD4 can interact directly with both DNMT1 and MLH1 leading to recruitment of all three components at sites of DNA damage (Ruzov et al., 2009). The co-localization of DNMT1, MBD4 and MLH1 suggests that they may participate in a cellular checkpoint that monitors potential DNA hypomethylation events by detecting the presence or absence of the maintenance methyltransferase Dnmt1, perhaps at or adjacent to the replication fork. The recruitment of these components in response to localized DNA damage suggests that can have a role in the cellular decision whether to repair the lesion or activate apoptosis (Ruzov et al., 2009). Decreases in DNMT1 protein levels in ES cells and normal differentiated human cells can result in mismatch repair (MMR) defects (Guo et al., 2004; Kim et al., 2004; Loughery et al., 2011; Wang and James Shen, 2004) In the hTERT-1604 fibroblast cell line, a normal diploid cell line immortalised by telomerase over-expression, depletion of DNMT1 is sufficient to cause MMR defects and increased mutation rates at a CA17 microsatellite. This is associated with decreases in MMR protein levels, including MBD4: following activation of the DNA damage response (DDR). Blocking the DDR, and in particular PARP over-activation, also increases survival of the DNMT1 knockdowns (Loughery et al., 2011).
zMbd4 has been proposed to be a candidate protein that is involved in DNA demethylation activity in Zebrafish (Rai et al., 2008). Rai and colleagues showed MBD4 removes G:T mismatch-specific thymines, resulting from deamination of 5-methylcytosine (5-meC) via cytidine deaminase family members (Activation Induced deaminase (AID) and Apolipo-protein B RNA-editing catalytic component (Apobec)). Interestingly, deamination activity by AID/Apobec did not occur unless MBD4 and/or other possible factor are present and/or activated, and a catalytically inactive hMBD4 derivative (D560A) stabilized the putative G:T intermediate and prevented rapid thymine removal (Rai et al., 2008). MBD4 was also reported to interact with Xenopus DNMT1 to promote p53 dependent apoptosis (Ruzov et al., 2009), and DNMT3b, which was is proposed to methylate cytosine and to deaminate 5-meC, relying on an inefficient deaminase activity (Kangaspeska et al., 2008; Metivier et al., 2008). Recent studies in mice have implicated thymine DNA-glycosylase (TDG) in active DNA demethylation, TDG interacts with the deaminase AID and the damage response protein GADD45a. These findings suggest a two-step mechanism for DNA demethylation in mammals, whereby 5-methylcytosine and 5-hydroxymethylcytosine are first deaminated by AID to thymine and 5-hydroxymethyluracil, respectively, followed by TDG-mediated thymine and 5-hydroxymethyluracil excision repair (Cortazar et al., 2011; Cortellino et al., 2011). It is possible that MBD4 may make a contribution to this process.
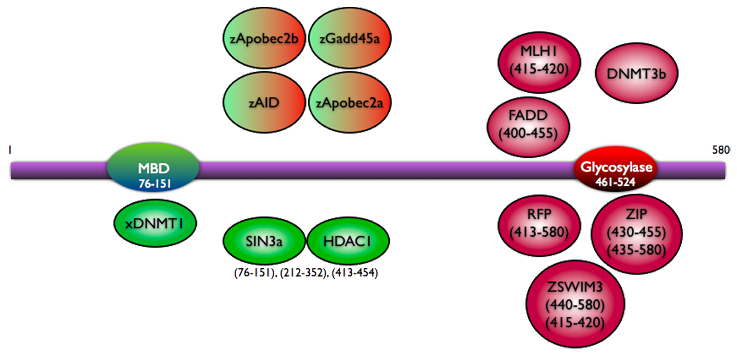
MBD4 interactions.
Mutations
Note
A C-terminal truncation of MBD4 containing its MBD domain was detected in the HCA7 cell line (colon cancer cell line) (Bader et al., 1999). Frequent MBD4 mutations were reported in MSI tumours in sporadic colon cancers and in hereditary non-polyposis colorectal cancer (HNPCC) tumours. The frequency of MBD4 mutations was correlated with MSH2/MLH1 germline mutations (Riccio et al., 1999). In humans, frame-shift mutation of MBD4 occurs in up to 43% of microsatellite unstable colon cancers. Recombinant truncated MBD4 (MBD4tru) inhibits glycosylase activities of normal MBD4 or Uracil DNA glycosylase in cell-free assays as a dominant negative effect. Over-expression of MBD4tru in Big Blue (lacI)-transfected, MSI human colorectal carcinoma cells doubled the mutation frequency, indicating that the modest dominant negative effect on DNA repair can occur in living cells in short-term experiments. Intriguingly, the whole mutation spectrum was increased, not only at CpG sites, suggesting that truncated MBD4 has a more widespread effect on genomic stability. This demonstration of a dominant negative effect may be of significance in tumour progression and acquisition of drug resistance (Abdel-Rahman et al., 2008; Bader et al., 2007).
Implicated in
Entity name
Intestinal tumorigenesis
Note
A reduced apoptotic response occurs in the small intestine of mbd4-/- mice in response to a variety of DNA-damaging agents, and increased tumorigenicity was observed on a background of mbd4-/- with the tumor-susceptible Apc min background in mice (Millar et al., 2002; Sansom et al., 2003; Wong et al., 2002). A frame-shift in a polyadenine tract due to an MMR defective background resulted in a mutated form of MBD4 that occurs naturally in human colon cancers (Abdel-Rahman et al., 2008). This leads to a premature truncation of the MBD4 protein, which has an intact MBD domain but lacks of the whole glycosylase domain. It has been suggested that this truncated form of MBD4 acts in a dominant negative way, competitively inhibiting normal glycosylase activity of wild type MBD4, and increases mutation frequency when overexpressed in cells (Bader et al., 2007). The frameshift mutations in MBD4 appear to be selected for in MSI tumour cells, as similar coding polyadenine repeats in other tested genes are rarely altered in this panel of 56 primary MSI tumours (Riccio et al., 1999).
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 18162445 | 2008 | Truncation of MBD4 predisposes to reciprocal chromosomal translocations and alters the response to therapeutic agents in colon cancer cells. | Abdel-Rahman WM et al |
| 18813943 | 2008 | Evolution of DNA-methylation machinery: DNA methyltransferases and methyl-DNA binding proteins in the amphioxus Branchiostoma floridae. | Albalat R et al |
| 10637515 | 1999 | Somatic frameshift mutations in the MBD4 gene of sporadic colon cancers with mismatch repair deficiency. | Bader S et al |
| 17285135 | 2007 | A human cancer-associated truncation of MBD4 causes dominant negative impairment of DNA repair in colon cancer cells. | Bader SA et al |
| 11121095 | 2001 | Methyl-CpG-binding proteins. Targeting specific gene repression. | Ballestar E et al |
| 10097147 | 1999 | MED1, a novel human methyl-CpG-binding endonuclease, interacts with DNA mismatch repair protein MLH1. | Bellacosa A et al |
| 21278727 | 2011 | Embryonic lethal phenotype reveals a function of TDG in maintaining epigenetic stability. | Cortázar D et al |
| 21722948 | 2011 | Thymine DNA glycosylase is essential for active DNA demethylation by linked deamination-base excision repair. | Cortellino S et al |
| 17049487 | 2006 | RET finger protein enhances MBD2- and MBD4-dependent transcriptional repression. | Fukushige S et al |
| 15215866 | 2004 | Mismatch repair genes identified using genetic screens in Blm-deficient embryonic stem cells. | Guo G et al |
| 12711219 | 2003 | The methyl-CpG binding domain and the evolving role of DNA methylation in animals. | Hendrich B et al |
| 18322535 | 2008 | Transient cyclical methylation of promoter DNA. | Kangaspeska S et al |
| 15509869 | 2004 | Dnmt1 deficiency leads to enhanced microsatellite instability in mouse embryonic stem cells. | Kim M et al |
| 16403636 | 2006 | Genomic DNA methylation: the mark and its mediators. | Klose RJ et al |
| 15899845 | 2005 | The thymine DNA glycosylase MBD4 represses transcription and is associated with methylated p16(INK4a) and hMLH1 genes. | Kondo E et al |
| 21636528 | 2011 | DNMT1 deficiency triggers mismatch repair defects in human cells through depletion of repair protein levels in a process involving the DNA damage response. | Loughery JE et al |
| 18322525 | 2008 | Cyclical DNA methylation of a transcriptionally active promoter. | Métivier R et al |
| 12130785 | 2002 | Enhanced CpG mutability and tumorigenesis in MBD4-deficient mice. | Millar CB et al |
| 19109892 | 2008 | DNA demethylation in zebrafish involves the coupling of a deaminase, a glycosylase, and gadd45. | Rai K et al |
| 10545939 | 1999 | The DNA repair gene MBD4 (MED1) is mutated in human carcinomas with microsatellite instability. | Riccio A et al |
| 19502488 | 2009 | MBD4 and MLH1 are required for apoptotic induction in xDNMT1-depleted embryos. | Ruzov A et al |
| 14562041 | 2003 | MBD4 deficiency reduces the apoptotic response to DNA-damaging agents in the murine small intestine. | Sansom OJ et al |
| 12702765 | 2003 | Fas-associated death domain protein interacts with methyl-CpG binding domain protein 4: a potential link between genome surveillance and apoptosis. | Screaton RA et al |
| 15378011 | 2004 | DNA methyltransferase Dnmt1 and mismatch repair. | Wang KY et al |
| 12417741 | 2002 | Mbd4 inactivation increases Cright-arrowT transition mutations and promotes gastrointestinal tumor formation. | Wong E et al |
Other Information
Locus ID:
NCBI: 8930
MIM: 603574
HGNC: 6919
Ensembl: ENSG00000129071
Variants:
dbSNP: 8930
ClinVar: 8930
TCGA: ENSG00000129071
COSMIC: MBD4
RNA/Proteins
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 38060262 | 2024 | Familial uveal melanoma and other tumors in 25 families with monoallelic germline MBD4 variants. | 0 |
| 38060262 | 2024 | Familial uveal melanoma and other tumors in 25 families with monoallelic germline MBD4 variants. | 0 |
| 37957685 | 2023 | MBD4 loss results in global reactivation of promoters and retroelements with low methylated CpG density. | 0 |
| 37957685 | 2023 | MBD4 loss results in global reactivation of promoters and retroelements with low methylated CpG density. | 0 |
| 35863105 | 2022 | MBD4 deficiency is predictive of response to immune checkpoint inhibitors in metastatic uveal melanoma patients. | 8 |
| 35863105 | 2022 | MBD4 deficiency is predictive of response to immune checkpoint inhibitors in metastatic uveal melanoma patients. | 8 |
| 32239153 | 2021 | Germline MBD4 Mutations and Predisposition to Uveal Melanoma. | 30 |
| 33871441 | 2021 | [Comparative Analysis of the Activity of the Polymorphic Variants of Human Uracil-DNA-Glycosylases SMUG1 and MBD4]. | 0 |
| 34107280 | 2021 | Structural Insights into the Mechanism of Base Excision by MBD4. | 6 |
| 32239153 | 2021 | Germline MBD4 Mutations and Predisposition to Uveal Melanoma. | 30 |
| 33871441 | 2021 | [Comparative Analysis of the Activity of the Polymorphic Variants of Human Uracil-DNA-Glycosylases SMUG1 and MBD4]. | 0 |
| 34107280 | 2021 | Structural Insights into the Mechanism of Base Excision by MBD4. | 6 |
| 32421892 | 2020 | Germline loss-of-function variants in MBD4 are rare in Finnish patients with uveal melanoma. | 5 |
| 32421892 | 2020 | Germline loss-of-function variants in MBD4 are rare in Finnish patients with uveal melanoma. | 5 |
| 31476572 | 2019 | Modification of the base excision repair enzyme MBD4 by the small ubiquitin-like molecule SUMO1. | 4 |
Citation
Huan X Meng ; Richard R Meehan
MBD4 (methyl-CpG binding domain protein 4)
Atlas Genet Cytogenet Oncol Haematol. 2011-07-01
Online version: http://atlasgeneticsoncology.org/gene/41312/mbd4
