PCNA (proliferating cell nuclear antigen)
2011-10-01 Ivaylo Stoimenov , Thomas Helleday AffiliationDepartment of Genetics Microbiology, Toxicology, Stockholm University, S-106 91 Stockholm, Sweden (IS, TH); Gray Institute for Radiation Oncology & Biology, University of Oxford, Oxford, OX3 7DQ, UK (TH)
DNA/RNA
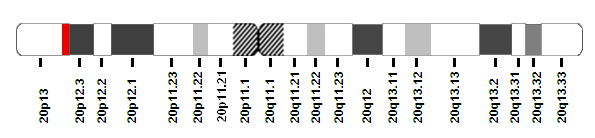
Description
Transcription
| NCBI Reference Sequence | Length (unspliced) | Length (spliced) | Exons | Protein | AA | |
| PCNA transcript variant 1 | NM_002592.2 | 11670 bp | 1355 bp | 7 | NP_002583.1 | 261 |
| PCNA transcript variant 2 | NM_182649.1 | 5049 bp | 1319 bp | 6 | NP_872590.1 | 261 |
PCNA transcript variant 1 is 1355 bp long after the completion of mRNA splicing. It has NCBI Reference Sequence code NM_002592.2 (NCBI). The PCNA transcript variant 1 has seven exons, six of which are contributing to the protein sequence. The first intron is relatively large in comparison with the other PCNA transcript variant. Following the splicing the length of the transcript is shortened to about 12% of that of the initial transcript. The translation starts from the middle of the 2nd exon and ends in the beginning of 7th exon. The product is a full length protein, designated as NP_002583.1 (NCBI), with 261 amino acids.
PCNA transcript variant 2 is 1319 bp long after the completion of mRNA splicing. It has NCBI Reference Sequence code NM_182649.1 (NCBI). The PCNA transcript variant 2 has six exons, which are contributing to the protein sequence. After the splicing the length of the transcript is shortened to about 26% of that of the initial transcript. Translation starts from the end of the 1st exon and ends in the beginning of 7th exon. The product is a full length protein, designated as NP_872590.1 (NCBI), with 261 amino acids.
Pseudogene
PCNAP1 and PCNAP2 - two pseudogenes in tandem on human chromosome 4 - q24 (Taniguchi et al., 1996).
There are several other possible pseudogenes:
LOC390102 on chromosome 11 - p15.1 (Webb et al., 1990),
LOC392454 on chromosome X - p11.3 (Ku et al., 1989; Webb et al., 1990).
Proteins

Description
Expression
Localisation
Function
Implicated in
According to OMIM and Human Locus Specific Mutation Databases there is no known disease, which is caused by mutation or loss of function of the PCNA protein.
The only implication of PCNA in human disease is as a prognostic or diagnostic marker, sometimes used together with other markers. The utilisation of PCNA as a marker is very much restricted to an illustration of proliferation potential and therefore cannot be specific for any disease. However, PCNA is indeed used as a prognostic and diagnostic marker in several human diseases in clinical practice, as shown below. The list is far from complete since any human disease associated with proliferation could utilise PCNA as a marker.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 9851246 | 1998 | Proliferating cell nuclear antigen (PCNA) immunolabeling as a prognostic factor in invasive ductal carcinoma of the breast in Taiwan. | Chu JS et al |
| 12040436 | 2002 | Expression profile of 11 proteins and their prognostic significance in patients with chronic lymphocytic leukemia (CLL). | Faderl S et al |
| 6600614 | 1983 | Clinical features of patients with antibodies directed against proliferating cell nuclear antigen. | Fritzler MJ et al |
| 9538167 | 1998 | Long-term prognostic value of PCNA labeling index in primary operable breast cancer. | Horiguchi J et al |
| 10482011 | 1999 | Immunohistochemical staining of proliferating cell nuclear antigen (PCNA) in malignant and nonmalignant skin diseases. | Kawahira K et al |
| 9711925 | 1998 | Prognostic implications of proliferating cell nuclear antigen (PCNA), AgNORs and P53 in non-Hodgkin's lymphomas. | Korkolopoulou P et al |
| 2569765 | 1989 | Human gene for proliferating cell nuclear antigen has pseudogenes and localizes to chromosome 20. | Ku DH et al |
| 11093818 | 2000 | Single nucleotide polymorphism analyses of the human proliferating cell nuclear antigen (pCNA) and flap endonuclease (FEN1) genes. | Ma X et al |
| 102692 | 1978 | Autoantibody to a nuclear antigen in proliferating cells. | Miyachi K et al |
| 17512402 | 2007 | PCNA, the maestro of the replication fork. | Moldovan GL et al |
| 15805117 | 2005 | Proliferating cell nuclear antigen (PCNA) may function as a double homotrimer complex in the mammalian cell. | Naryzhny SN et al |
| 18726183 | 2008 | Proliferating cell nuclear antigen: a proteomics view. | Naryzhny SN et al |
| 18304001 | 2008 | The biochemistry of somatic hypermutation. | Peled JU et al |
| 2882424 | 1987 | Functional identity of proliferating cell nuclear antigen and a DNA polymerase-delta auxiliary protein. | Prelich G et al |
| 18854411 | 2008 | Ubiquitylated PCNA plays a role in somatic hypermutation and class-switch recombination and is required for meiotic progression. | Roa S et al |
| 19442257 | 2009 | PCNA on the crossroad of cancer. | Stoimenov I et al |
| 3745189 | 1986 | An auxiliary protein for DNA polymerase-delta from fetal calf thymus. | Tan CK et al |
| 8995762 | 1996 | Cloning, sequencing, and chromosomal localization of two tandemly arranged human pseudogenes for the proliferating cell nuclear antigen (PCNA). | Taniguchi Y et al |
| 2565339 | 1989 | Structure of the human gene for the proliferating cell nuclear antigen. | Travali S et al |
| 1979311 | 1990 | Localization of the gene for human proliferating nuclear antigen/cyclin by in situ hybridization. | Webb G et al |
| 1350226 | 1992 | Prognostic value of proliferating cell nuclear antigen expression in chronic lymphoid leukemia. | del Giglio A et al |
Other Information
Locus ID:
NCBI: 5111
MIM: 176740
HGNC: 8729
Ensembl: ENSG00000132646
Variants:
dbSNP: 5111
ClinVar: 5111
TCGA: ENSG00000132646
COSMIC: PCNA
RNA/Proteins
| Gene ID | Transcript ID | Uniprot |
|---|---|---|
| ENSG00000132646 | ENST00000379143 | P12004 |
| ENSG00000132646 | ENST00000379160 | P12004 |
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 37824822 | 2024 | G-CSF reshapes the cytosolic PCNA scaffold and modulates glycolysis in neutrophils. | 1 |
| 38261971 | 2024 | Checkpoint activation by Spd1: a competition-based system relying on tandem disordered PCNA binding motifs. | 1 |
| 38280406 | 2024 | Squalene monooxygenase facilitates bladder cancer development in part by regulating PCNA. | 0 |
| 38387257 | 2024 | Loci on chromosome 20 interact with rs16969968 to influence cigarettes per day in European ancestry individuals. | 0 |
| 38648458 | 2024 | Expression of CD44, PCNA and E-cadherin in pterygium tissues. | 0 |
| 38669181 | 2024 | Cryo-EM reveals a nearly complete PCNA loading process and unique features of the human alternative clamp loader CTF18-RFC. | 0 |
| 38687509 | 2024 | EIF3B stabilizes PCNA by counteracting SYVN1-mediated ubiquitination to serve as a promotor in cholangiocarcinoma. | 0 |
| 38713100 | 2024 | DNAJC24 acts directly with PCNA and promotes malignant progression of LUAD by activating phosphorylation of AKT. | 0 |
| 37824822 | 2024 | G-CSF reshapes the cytosolic PCNA scaffold and modulates glycolysis in neutrophils. | 1 |
| 38261971 | 2024 | Checkpoint activation by Spd1: a competition-based system relying on tandem disordered PCNA binding motifs. | 1 |
| 38280406 | 2024 | Squalene monooxygenase facilitates bladder cancer development in part by regulating PCNA. | 0 |
| 38387257 | 2024 | Loci on chromosome 20 interact with rs16969968 to influence cigarettes per day in European ancestry individuals. | 0 |
| 38648458 | 2024 | Expression of CD44, PCNA and E-cadherin in pterygium tissues. | 0 |
| 38669181 | 2024 | Cryo-EM reveals a nearly complete PCNA loading process and unique features of the human alternative clamp loader CTF18-RFC. | 0 |
| 38687509 | 2024 | EIF3B stabilizes PCNA by counteracting SYVN1-mediated ubiquitination to serve as a promotor in cholangiocarcinoma. | 0 |
Citation
Ivaylo Stoimenov ; Thomas Helleday
PCNA (proliferating cell nuclear antigen)
Atlas Genet Cytogenet Oncol Haematol. 2011-10-01
Online version: http://atlasgeneticsoncology.org/gene/41670/pcna-(proliferating-cell-nuclear-antigen)
