CHD6 (chromodomain helicase DNA binding protein 6)
2014-08-01 Melissa Lathrop , C Harker Rhodes , Steve Fiering AffiliationDartmouth-Hitchcock Medical Center, Lebanon, New Hampshire, USA
Identity
Abstract
CHD6 is a chromatin remodeling protein characterized to play a role in transcriptional repression of genes and viruses. It occurs in a nuclear location as a component of a larger complex which associates with RNA Pol II. Mutations in CHD6 are associated with motor coordination defects, and development of cancers following substitutions and translocations.
DNA/RNA
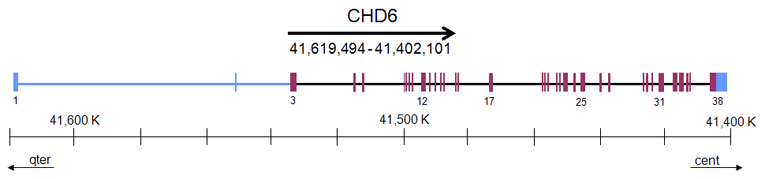
Description
Transcription
Pseudogene
Proteins
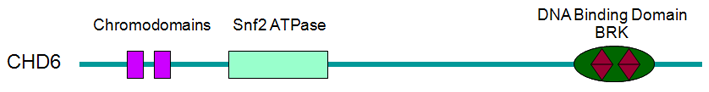
Description
Expression
Localisation
Function
Homology
Mutations
Note
Implicated in
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 21899694 | 2011 | CHD6 chromatin remodeler is a negative modulator of influenza virus replication that relocates to inactive chromatin upon infection. | Alfonso R et al |
| 23408615 | 2013 | CHD6, a cellular repressor of influenza virus replication, is degraded in human alveolar epithelial cells and mice lungs during infection. | Alfonso R et al |
| 24694993 | 2014 | Integrated analysis of copy number variation and genome-wide expression profiling in colorectal cancer tissues. | Ali Hassan NZ et al |
| 17713580 | 2007 | How many remodelers does it take to make a brain? Diverse and cooperative roles of ATP-dependent chromatin-remodeling complexes in development. | Brown E et al |
| 24844365 | 2015 | A novel translocation (6;20)(q13;q12) in acute myeloid leukemia likely results in LMBRD1-CHD6 fusion. | Douet-Guilbert N et al |
| 20631145 | 2010 | Interaction of the papillomavirus E8--E2C protein with the cellular CHD6 protein contributes to transcriptional repression. | Fertey J et al |
| 21473668 | 2012 | Genome-wide association study of motor coordination problems in ADHD identifies genes for brain and muscle function. | Fliers EA et al |
| 21822268 | 2011 | Frequent mutations of chromatin remodeling genes in transitional cell carcinoma of the bladder. | Gui Y et al |
| 20095039 | 2010 | Characterization of chromosome arm 20q abnormalities in myeloid malignancies using genome-wide single nucleotide polymorphism array analysis. | Huh J et al |
| 20111866 | 2010 | Deletion of the Chd6 exon 12 affects motor coordination. | Lathrop MJ et al |
| 17027977 | 2006 | CHD6 is a DNA-dependent ATPase and localizes at nuclear sites of mRNA synthesis. | Lutz T et al |
| 17350655 | 2007 | The Chd family of chromatin remodelers. | Marfella CG et al |
| 24755471 | 2014 | Colorectal cancer cell lines are representative models of the main molecular subtypes of primary cancer. | Mouradov D et al |
| 16314513 | 2005 | The carboxy-terminal Neh3 domain of Nrf2 is required for transcriptional activation. | Nioi P et al |
| 24068064 | 2013 | Isolated deletion of the long arm of chromosome 20 [del(20q12)] in myelodysplastic syndrome: a case report and literature review. | Padhi S et al |
| 11889561 | 2002 | CHD5 defines a new subfamily of chromodomain-SWI2/SNF2-like helicases. | Schuster EF et al |
| 23954449 | 2013 | CHD chromatin remodelling enzymes and the DNA damage response. | Stanley FK et al |
| 16638715 | 2006 | Identification of differentially transcribed genes in human lymphoblastoid cells irradiated with 0.5 Gy of gamma-ray and the involvement of low dose radiation inducible CHD6 gene in cell proliferation and radiosensitivity. | Wang HP et al |
| 21086493 | 2010 | Characterization of a de novo balanced t(4;20)(q33;q12) translocation in a patient with mental retardation. | Yamada K et al |
Other Information
Locus ID:
NCBI: 84181
MIM: 616114
HGNC: 19057
Ensembl: ENSG00000124177
Variants:
dbSNP: 84181
ClinVar: 84181
TCGA: ENSG00000124177
COSMIC: CHD6
RNA/Proteins
Expression (GTEx)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 36408932 | 2022 | CHD6 promotes broad nucleosome eviction for transcriptional activation in prostate cancer cells. | 2 |
| 36408932 | 2022 | CHD6 promotes broad nucleosome eviction for transcriptional activation in prostate cancer cells. | 2 |
| 34021162 | 2021 | Overarching control of autophagy and DNA damage response by CHD6 revealed by modeling a rare human pathology. | 12 |
| 34021162 | 2021 | Overarching control of autophagy and DNA damage response by CHD6 revealed by modeling a rare human pathology. | 12 |
| 30651562 | 2019 | The CHD6 chromatin remodeler is an oxidative DNA damage response factor. | 33 |
| 30651562 | 2019 | The CHD6 chromatin remodeler is an oxidative DNA damage response factor. | 33 |
| 28533432 | 2017 | The ATP-dependent chromatin remodeling enzymes CHD6, CHD7, and CHD8 exhibit distinct nucleosome binding and remodeling activities. | 38 |
| 28533432 | 2017 | The ATP-dependent chromatin remodeling enzymes CHD6, CHD7, and CHD8 exhibit distinct nucleosome binding and remodeling activities. | 38 |
| 25631877 | 2015 | CHD6 regulates the topological arrangement of the CFTR locus. | 11 |
| 25631877 | 2015 | CHD6 regulates the topological arrangement of the CFTR locus. | 11 |
| 25631877 | 2015 | CHD6 regulates the topological arrangement of the CFTR locus. | 11 |
| 25631877 | 2015 | CHD6 regulates the topological arrangement of the CFTR locus. | 11 |
| 23408615 | 2013 | CHD6, a cellular repressor of influenza virus replication, is degraded in human alveolar epithelial cells and mice lungs during infection. | 9 |
| 23408615 | 2013 | CHD6, a cellular repressor of influenza virus replication, is degraded in human alveolar epithelial cells and mice lungs during infection. | 9 |
| 21899694 | 2011 | CHD6 chromatin remodeler is a negative modulator of influenza virus replication that relocates to inactive chromatin upon infection. | 27 |
Citation
Melissa Lathrop ; C Harker Rhodes ; Steve Fiering
CHD6 (chromodomain helicase DNA binding protein 6)
Atlas Genet Cytogenet Oncol Haematol. 2014-08-01
Online version: http://atlasgeneticsoncology.org/gene/43211/chd6-(chromodomain-helicase-dna-binding-protein-6)
