NUP214/ABL1 fusion gene on amplified episomes
2012-06-01 Nathalie Nadal Affiliation1.Laboratoire dhematologie, Pavillon de Biologie, CHU Hopital Nord, 42055 St Etienne Cedex 2, France
Clinics and Pathology
Disease
Note
Not seen in B-cell ALL or other malignant diseases. , The main hypothesis is that genomic amplification is a dynamic process. Molecular chronology of genomic amplification has been schematically described as follows. The first step is the production of submicroscopic, acentric, circular, extrachromosomal DNA molecules which replicate autonomously, called episomes. These DNA molecules are made of amplified genes. 2 mechanisms for the formation of episomes have been proposed: Conservative which preserves the original DNA sequence at the native chromosomal locus and non conservative which leads to the deletion of the original sequence at the native locus. The second step corresponds to an increase in copy number resulting from unequal mitotic segregation and an increase in size. They enlarge over time to form progressively heterogeneously sized structures, microscopically visible, called double minutes (dmin). In a later step they may integrate into chromosomes to generate intrachromosomally amplified structures (HSR). In some cases dmins or HSRs may form directly without precursors.
Phenotype stem cell origin
Immature T-cell leukemia (CD3+, CD2+ and CD7+).
Epidemiology
Clinics
No major clinical differences between NUP214-ABL negative and positive T-ALL.
Cytology
Lymphoblasts.
Treatment
NUP214-ABL1 cells are sensitive to tyrosine kinase inhibitors (ITK). Targeting therapies may improve outcome of patients with T-ALL expressing NUP214-ABL1 but the clinical experience is, yet, too limited to conclude.
Prognosis
Most data, but not all, suggests that NUP214-ABL1 fusion gene amplification in T-ALL is associated with poor outcome.
Note
Mechanism of gene amplification
Cytogenetics
Cytogenetics morphological
Not detectable by conventional cytogenetics. Cryptic, no dmin.
Cytogenetics molecular
FISH using commercially available ABL1 probe shows multiple extrachromosomal sites on metaphases and multiple signals in interphase nuclei. The extrachromosomal amplification of ABL1 appears to be pathognomonique for the presence of NUP214-ABL1 fusion in T-ALL.There may be a corresponding deletion of the ABL1 probe on one of the chromosomes 9 (see note above concerning mechanisms of gene amplification).
Probes
ABL1 probe.
Additional anomalies
None. In apparently normal karyotype or with variable additional abnormalities.
Genes Involved and Proteins
Gene name
ABL1 (v-abl Abelson murine leukemia viral oncogene homolog 1)
Location
9q34.12
Dna rna description
Alternate splicing. mRNA of 6 and 7 kb. , 7.5 kb mRNA.
Protein description
Protein 145 kDa; Localization: nuclear and cytoplasmic; Tyrosine kinase; Ubiquitously expressed. ABL1 modulates T-cell development and plays a role in cytoskeletal remodelling processes in T-cells.
Gene name
NUP214 (nucleoporin 214kDa)
Location
9q34.13
Note
More telomeric than ABL1.
Dna rna description
Other names: CAN, CAIN, Nucleoporin.
Protein description
Component of the Nuclear Pore Complex. 214 kDa; 2 dimerization domains (2 leucine zippers) and a repeated motif; forms homodimers. Mediate nucleocytoplasmic transport. Localisation: nuclear membrane; cytoplasmic face.
Result of the Chromosomal Anomaly
Note
NUP214 is recognized as being the second most prevalent fusion gene involving ABL1.
Description
Molecular analyses delineated the amplicon as a 500 kb region from chromosome band 9q34 containing the genes ABL1, LAMC3 and NUP214. The genomic region from ABL1 to NUP214 circularizes to generate the NUP214-ABL1 fusion gene = New mechanism for generation of a fusion gene. The breakpoint within ABL1 occurs in intron 1 in most cases, in intron 2 in other cases (coincides with ABL1 breakpoint in the Philadelphia chromosome). Whilst the breakpoint in NUP214 is variable (ranging from intron 23 to intron 34).
Transcript
NUP214-ABL1 fusion gene.
Detection protocole
RT-PCR of the fusion transcript.
Description
The NUP214-ABL transcript encodes a 239-333 kDa protein which includes the coiled-coil domain (dimerisation motifs necessary for tyrosine kinase activation and neoplastic transformation) of NUP214 and the tyrosine kinase domain of ABL1.
Description
NUP214-ABL protein is a constitutively activated tyrosine kinase most likely implicated in the pathogenesis of T-ALL in a similar mechanism of action as for BCR-ABL as it activates similar pathways and for its sensitivity to ITK. However NUP214-ABL protein is less potent and requires amplification for neoplastic transformation.
Oncogenesis
NUP214 is also involved in the translocation t(6;9) seen in myeloid malignancies which results in the fusion gene DEK-NUP214. Unlike NUP214-ABL where the N-terminal region of NUP214 is retained, in DEK-NUP214, it is the C-terminal region of NUP214 which is present. The mode of leukemogenesis of DEK-NUP214 is thought to be interference with nucleocytoplasmic transport processes. It is unknown whether NUP214-ABL acts in the same way.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 15674415 | 2005 | NUP214-ABL1 amplification in t(5;14)/HOX11L2-positive ALL present with several forms and may have a prognostic significance. | Ballerini P et al |
| 15057249 | 2004 | Amplification of the ABL gene in T-cell acute lymphoblastic leukemia. | Barber KE et al |
| 16873673 | 2006 | NUP214-ABL1 in adult T-ALL: the GMALL study group experience. | Burmeister T et al |
| 21435002 | 2011 | ABL1 fusion genes in hematological malignancies: a review. | De Braekeleer E et al |
| 15713800 | 2005 | Fusion of EML1 to ABL1 in T-cell acute lymphoblastic leukemia with cryptic t(9;14)(q34;q32). | De Keersmaecker K et al |
| 15361874 | 2004 | Fusion of NUP214 to ABL1 on amplified episomes in T-cell acute lymphoblastic leukemia. | Graux C et al |
| 18401417 | 2008 | Activity of tyrosine kinase inhibitors against human NUP214-ABL1-positive T cell malignancies. | Quintás-Cardama A et al |
| 16015385 | 2005 | Fusion of NUP214 to ABL1 on amplified episomes in T-ALL--implications for treatment. | Stergianou K et al |
| 2647287 | 1989 | The importance of circular DNA in mammalian gene amplification. | Wahl GM et al |
Summary
Note
Episomes are submicroscopic extrachromosomal structures.
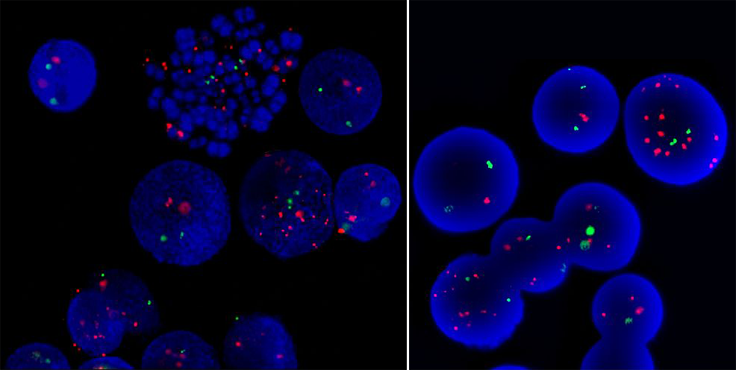
Episomal amplification of ABL detected by FISH with the commercial probe LSI BCR-ABL ES.
Citation
Nathalie Nadal
NUP214/ABL1 fusion gene on amplified episomes
Atlas Genet Cytogenet Oncol Haematol. 2012-06-01
Online version: http://atlasgeneticsoncology.org/haematological/1397/nup214-abl1-fusion-gene-on-amplified-episomes
Historical Card
2005-09-01 NUP214/ABL1 fusion gene on amplified episomes by Nathalie Nadal Affiliation
Laboratoire dhematologie, Pavillon de Biologie, CHU Hopital Nord, 42055 St Etienne Cedex 2, France
