t(10;11)(q22;q23) KMT2A/TET1
2014-06-01 Antoine Ittel Affiliation1.Laboratoire de Cytogenetique hematologique, CHU Hautepierre, Strasbourg, France
Clinics and Pathology
Disease
Acute myeloid leukemia (AML), B-cell precursor acute lymphoblastic leukemia (ALL), T-cell lymphoblastic lymphoma
Phenotype stem cell origin
Described in subtypes AML-M2, -M4 and -M5. Cell lineage dysplasia may be associated. Described also in 2 B-ALL and one case of T-cell lymphoblastic lymphoma with a subsequent transformation to AML.
Epidemiology
Described in 14 cases in the literature, mainly adults and 2 children (8 men, 5 women, 1 unknown) median age of 39 years (range 1 month to 67 years).
Frequency estimated at 0.3 % of MLL-rearranged acute leukemia cases (5 out 1590) (Lee et al., 2013).
Frequency estimated at 0.3 % of MLL-rearranged acute leukemia cases (5 out 1590) (Lee et al., 2013).
Prognosis
Undetermined, possibly intermediate. On the 8 of 14 patients with available clinical outcomes, 7 had a complete remission. 6 patients died at an average of 16 months after initial diagnosis.
Cytogenetics
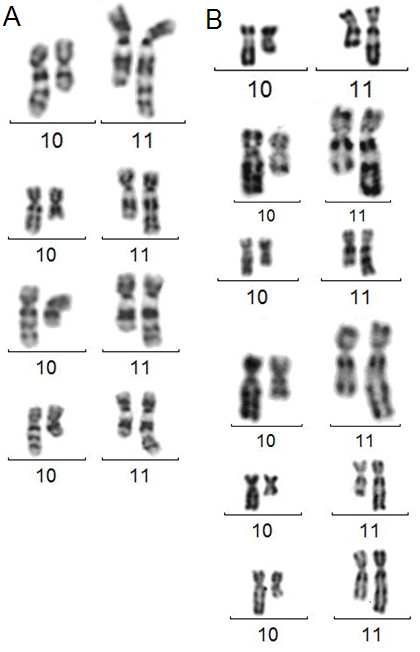
A. t(10;11)(q22;q23) in GTG banding. B. The same with RHG banding.
Cytogenetics morphological
Easy to detect, evident 10q- and 11q+ derivatives.
Cytogenetics molecular
Commercial dual color MLL/KMT2A FISH probes are splitted by the translocation. 10q22 breakpoint may be detected with RP11-119F7 BAC probe or using RP11-9E13 and RP11-314J18 BACs (separation of the two BACs with the t(10;11)(q22;q23)).

A. FISH on metaphase (inverted DAPI) using RP11-9E13 and RP11-314J18 BACs showing the separation of the two probes in the t(10;11)(q22;q23). B. Focus on chromosomes 10 and 11 and derivative chromosomes (inverted DAPI) from the t(10;11) with the same BACs.
Additional anomalies
5/11 caryotypes available have the t(10;11) as sole chromosomal abnormality. 4/11 have one more chromosomal abnormality (+21, del(6)p(21), del(5)(q15), add(13)(p11) or der(6)t(6;?9)(p22;?q21) in 2 different clones with the t(10;11)); one case was hyperdiploid with 51 chromosomes; one case had complex caryotype.
Genes Involved and Proteins
Gene name
KMT2A (myeloid/lymphoid or mixed lineage leukemia)
Location
11q23.3
Note
KMT2A is also called MLL (mixed-lineage leukemia or myeloid-lymphoid leukaemia), ALL-1 or HRX.
Dna rna description
36 exons, multiple transcripts 13-15 kb.
Protein description
430 kDA, contains two DNA binding motifs (a AT hook and a CXXC domain), a DNA methyl transferase motif, a bromodomain; transcriptional regulatory factor involved in maintenance of Hox gene expression during embryogenesis and during the process of haematopoietic progenitors expansion and differentiation.
Gene name
TET1 (tet methylcytosine dioxygenase 1)
Location
10q21.3
Note
TET1 is also called LCX (leukemia-associated protein with a CXXC domain) or CXXC6 (CXXC finger 6).
Dna rna description
8497 bp representing the whole coding sequence. Contains 12 exons. Contains 3 bipartite nuclear localization sites, 1 alpha helice coiled-coil region and 1 cysteine rich domain with high level homology with a CXXC DNA binding site.
Protein description
Predicted size of 2136 amino acids, expression restricted to some fetal tissues, mainly lung, heart and brain; not expressed in hematopoietic tissues, except in spleen; unknown function.
Protein description
TET family enzymes convert 5-methylcytosine (5mC) to 5-hydroxymethylcytosine (5hmC). Key role in active DNA demethylation. TET1 and TET2: key enzymes responsible for the presence of 5hmC in mouse embryonic stem cells (ESCs). TET1: regulates the lineage differentiation potential of ESCs. TET1 interacts physically with NANOG, synergistically enhancing the efficiency of NANOG in somatic cell reprogramming. NANOG/TET1 co-occupy genomic loci of genes associated with both maintenance of pluripotency and lineage commitment in embryonic stem cells, and may deposit 5hmC to target genes before the establishment of pluripotency. Taken together, these observations suggest a possible mechanism for the lineage switch observed in one of the 14 cases. (Ittel et al., 2013).
Protein description
TET1 significantly up-regulated in MLL-rearranged leukemia. TET1: direct target gene of MLL-fusion proteins. MLL fusions bind to the promoter region of TET1 and promote its expression directly in both human and mouse hematopoietic stem/progenitor cells, cumulating in a global increase of 5hmC. Briefly, MLL-fusion proteins bind directly to the Tet1 locus. Consequently this promote its expression and the increased expression of Tet1 (and the corresponding global increase of 5hmC) cooperates with MLL fusions in orchestrating the transcriptional activation of their cotargets. The main critical oncogenic cotargets described are HOXA9, MEIS1 and PBX3. The Hoxa9/Meis1/Pbx3 signaling cascade promotes cell proliferation and inhibits apoptosis/cell differentiation, thereby leading to cell transformation and leukemogenesis. (Huang et al., 2013).
Result of the Chromosomal Anomaly
Description
Transcripts from the 5 MLL-LCX 3 fusion gene on der(11) are expressed; transcripts from the 5 LCX-MLL 3 counterpart are not detected.
Breakpoints in the MLL gene are located between intron 6 and exon 11.
All genomic breakpoints within the TET1 gene were identified in an approximately 17 kb genomic region flanked by TET1 exons 8 and 12. Most characterized breakpoints (5 of 7 cases) were mapped to intron 8.Predicted molecular weight of 204.4 kDa.
Breakpoints in the MLL gene are located between intron 6 and exon 11.
All genomic breakpoints within the TET1 gene were identified in an approximately 17 kb genomic region flanked by TET1 exons 8 and 12. Most characterized breakpoints (5 of 7 cases) were mapped to intron 8.Predicted molecular weight of 204.4 kDa.
Oncogenesis
Unknown; the alpha helice coiled-coil region retained at the COOH extremity might be involved in the leukemogenesis.
Highly cited references
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 29948955 | 2018 | Correction to: Dysplastic features seen in a patient with acute myeloid leukemia harboring the KMT2A-TET1 fusion gene. | 0 |
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 9973924 | 1999 | Involvement of MLL gene in a t(10;11)(q22;q23) and a t(8;11)(q24;q23) identified by fluorescence in situ hybridization. | Aventín A et al |
| 23818607 | 2013 | TET1 plays an essential oncogenic role in MLL-rearranged leukemia. | Huang H et al |
| 24323992 | 2013 | First description of the t(10;11)(q22;q23)/MLL-TET1 translocation in a T-cell lymphoblastic lymphoma, with subsequent lineage switch to acute myelomonocytic myeloid leukemia. | Ittel A et al |
| 15375572 | 2004 | Identification and characterization of human CXXC10 gene in silico. | Katoh M et al |
| 23100278 | 2013 | Genomic breakpoints and clinical features of MLL-TET1 rearrangement in acute leukemias. | Lee SG et al |
| 12646957 | 2003 | TET1, a member of a novel protein family, is fused to MLL in acute myeloid leukemia containing the t(10;11)(q22;q23). | Lorsbach RB et al |
| 12124344 | 2002 | LCX, leukemia-associated protein with a CXXC domain, is fused to MLL in acute myeloid leukemia with trilineage dysplasia having t(10;11)(q22;q23). | Ono R et al |
Summary
Fusion gene
KMT2A/TET1 KMT2A (11q23.3) TET1 (10q21.3) TIC
Citation
Antoine Ittel
t(10;11)(q22;q23) KMT2A/TET1
Atlas Genet Cytogenet Oncol Haematol. 2014-06-01
Online version: http://atlasgeneticsoncology.org/haematological/1410/t(10;11)(q22;q23)-kmt2a-tet1
