CTNND1 (catenin (cadherin-associated protein), delta 1)
2010-05-01 Michael R Dohn , Albert B Reynolds AffiliationDepartment of Cancer Biology, Vanderbilt University, Nashville, TN, USA
Identity
HGNC
LOCATION
11q12.1
LOCUSID
ALIAS
BCDS2,CAS,CTNND,P120CAS,P120CTN,p120,p120(CAS),p120(CTN)
FUSION GENES
DNA/RNA
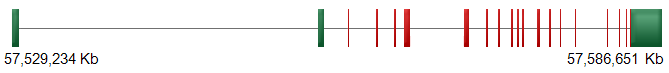
CTNND1/p120 gene structure. DNA of CTNND1 gene (NM_001085458.1) composed of 19 coding exons (red). Untranslated regions are depicted in green.
Description
DNA contains 57419 bp composed of 21 exons, 19 of which are coding exons.
Transcription
Alternative transcription produces several distinct mRNAs ranging from 5676 bp to 6329 bp, transcribed in centromeric to telomeric orientation, with open reading frames ranging from 2802 bp to 2904 bp.
Pseudogene
None.
Proteins

CTNND1/p120 protein structure. p120 contains four translational start sites (1-4, arrows) and three alternatively transcribed exons A, B, and C (red boxes). Isoform-1 contains an amino-terminal coiled coil domain (CC), and a phosphorylation domain (PD) is present in all but isoform-4. All isoforms contain a central Armadillo repeat domain (gray boxes).
Description
This gene encodes a member of the Armadillo (Arm) family of proteins, which function in cell-cell adhesion and signal transduction, and is the prototype for a subfamily of proteins that includes p120-catenin (hereafter p120), delta-catenin, p0071, and plakophilins 1-3 (PKP1, PKP2, PKP3). CTNND1/p120 has four distinct translation start codons (isoforms 1-4) and three alternatively spliced exons (A, B, and C), potentially resulting in 32 isoforms as products of alternative splicing. A coiled-coil domain is present in the extreme amino-terminus of the long-isoform (isoform-1), and an amino terminal phosphorylation domain is present in all but the shortest isoform (isoform-4). All p120 isoforms contain a central Arm domain consisting of nine Arm repeats. p120 has been shown to be post-translationally modified by phosphorylation on tyrosine, serine, and threonine residues, as well as by cleavage by the protease calpain. Phosphorylation of p120 may be regulated in part via interactions with non-receptor tyrosine kinases, (including Fer, Fyn, and Yes) and protein tyrosine phosphatases (including PTPmu, SHP-1, and DEP-1).
Expression
p120 isoforms have been detected in all tissues examined, although cells of epithelial origin tend to predominantly express isoform 3, whereas motile, fibroblastoid cells often express isoform 1.
Localisation
Cell-cell junctions (bound to cadherin molecules), cytoplasm, and nucleus.
Function
CTNND1/p120 was originally identified as a substrate for the Src oncogene and for various receptor tyrosine kinases, though its most prominent role is in stabilizing adherens junctions. Via its Arm domain, p120 binds to the juxtamembrane region of the cytoplasmic tails of type I and II cadherins. Upon removal of p120, by either gene knock-out or shRNA-mediated knock-down, adherens junction components, namely cadherin, beta-catenin, and alpha-catenin, are internalized and rapidly degraded. p120 also likely participates in the rearrangement of the actin cytoskeleton via regulation of Rho family GTPases, and p120 may also influence gene expression via its interaction in the nucleus and cytoplasm with the transcriptional repressor Kaiso.
Homology
Pan troglodytes - CTNND1; Canis lupus familiaris - CTNND1; Bos taurus - CTNND1; Mus musculus - Ctnnd1; Rattus norvegicus - Ctnnd1; Gallus gallus - RCJMB04_21j12; Danio rerio - LOC556726.
Mutations
Note
While p120 is frequently decreased in a wide variety of cancers, the only mutated p120 identified to date is found in colonic SW48 cells. In these cells, a heterozygous nonsense mutation in exon 7 at nucleotide 1908 (C to T) yields a premature stop codon that truncates the protein in the third ARM repeat. Several other cDNA abnormalities, including deletion of exon 17, retention of intron 19, or retention of both introns 19 and 20, were also identified in both p120 alleles in these cells, resulting in premature stop codons that eliminate the normal p120 COOH terminus.
Implicated in
Entity name
Various cancers
Note
p120s ability to stabilize E-cadherin, a tumor suppressor, suggests that p120 may play a role in tumor and/or metastasis suppression. Indeed, p120 is often lost, downregulated, or mislocalized in human colon, breast, prostate, lung, and other carcinomas. Total p120 knockout is embryonically lethal in mice, but conditional knockout studies have shown that p120 loss in the salivary gland induces dysplasia indistinguishable from high-grade intraepithelial neoplasia, and p120 ablation in the epidermis induces hyperproliferation and inflammation. However, recent studies have also identified a requirement for p120 in anchorage-independent growth of both tumor cell lines and oncogene-transformed cell lines. Moreover, a role for p120 and its binding partner Kaiso in Wnt signaling, which is often hyperactive in colon cancer, is becoming increasingly evident.
Entity name
Breast cancer
Note
cDNA microarray analysis revealed a downregulation of p120 in invasive breast cancers compared to their respective benign control breast tissues, and immunohistochemical analyses of tissue samples from 341 invasive breast cancer patients found that p120 downregulation predicted a 3.1-fold risk in breast cancer death.
Additionally, translation initiation factor eIF4GI-enhanced translation of IRES-containing p120 mRNAs has been shown to promote tumor emboli formation in inflammatory breast cancer.
Additionally, translation initiation factor eIF4GI-enhanced translation of IRES-containing p120 mRNAs has been shown to promote tumor emboli formation in inflammatory breast cancer.
Entity name
Lung cancer
Note
A study of 138 patients with non-small cell lung cancer found that p120 membrane localization was reduced and ectopic cytoplasmic expression was increased in lung cancer tissues. Abnormal p120 localization correlated with TNM stage and lymph node metastasis.
Entity name
Skin cancer
Note
p120 was found to be altered in squamous cell carcinomas in humans, and conditional ablation of p120 in the epidermis in mice led to progressive development of skin neoplasias.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 10321838 | 1999 | Human p120ctn catenin: tissue-specific expression of isoforms and molecular interactions with BP180/type XVII collagen. | Aho S et al |
| 10980705 | 2000 | Inhibition of RhoA by p120 catenin. | Anastasiadis PZ et al |
| 18950621 | 2009 | PDGF receptor activation induces p120-catenin phosphorylation at serine 879 via a PKCalpha-dependent pathway. | Brown MV et al |
| 15663951 | 2005 | Structure of the armadillo repeat domain of plakophilin 1. | Choi HJ et al |
| 10207085 | 1999 | The catenin p120(ctn) interacts with Kaiso, a novel BTB/POZ domain zinc finger transcription factor. | Daniel JM et al |
| 14610055 | 2003 | A core function for p120-catenin in cadherin turnover. | Davis MA et al |
| 16399075 | 2006 | Blocked acinar development, E-cadherin reduction, and intraepithelial neoplasia upon ablation of p120-catenin in the mouse salivary gland. | Davis MA et al |
| 19188496 | 2009 | An essential role for p120-catenin in Src- and Rac1-mediated anchorage-independent cell growth. | Dohn MR et al |
| 1851549 | 1991 | PDGF, CSF-1, and EGF induce tyrosine phosphorylation of p120, a pp60src transformation-associated substrate. | Downing JR et al |
| 11171375 | 2001 | p120 catenin affects cell motility via modulation of activity of Rho-family GTPases: a link between cell-cell contact formation and regulation of cell locomotion. | Grosheva I et al |
| 12370829 | 2002 | The transmembrane receptor protein tyrosine phosphatase DEP1 interacts with p120(ctn). | Holsinger LJ et al |
| 12427869 | 2002 | A novel role for p120 catenin in E-cadherin function. | Ireton RC et al |
| 1703631 | 1991 | Tyrosine phosphorylation of a 120-kilodalton pp60src substrate upon epidermal growth factor and platelet-derived growth factor receptor stimulation and in polyomavirus middle-T-antigen-transformed cells. | Kanner SB et al |
| 10835420 | 2000 | The protein-tyrosine phosphatase SHP-1 binds to and dephosphorylates p120 catenin. | Keilhack H et al |
| 9653641 | 1998 | Molecular cloning of the human p120ctn catenin gene (CTNND1): expression of multiple alternatively spliced isoforms. | Keirsebilck A et al |
| 15543138 | 2004 | Non-canonical Wnt signals are modulated by the Kaiso transcriptional repressor and p120-catenin. | Kim SW et al |
| 19162367 | 2009 | Abnormal expression of p120-catenin, E-cadherin, and small GTPases is significantly associated with malignant phenotype of human lung cancer. | Liu Y et al |
| 14996911 | 2004 | EGFR signaling to p120-catenin through phosphorylation at Y228. | Mariner DJ et al |
| 10931868 | 2000 | p120 catenin regulates the actin cytoskeleton via Rho family GTPases. | Noren NK et al |
| 17196166 | 2007 | Ischemia promotes calpain-mediated degradation of p120-catenin in SH-SY5Y cells. | Ohno H et al |
| 17084360 | 2006 | Frodo links Dishevelled to the p120-catenin/Kaiso pathway: distinct catenin subfamilies promote Wnt signals. | Park JI et al |
| 15935774 | 2005 | Kaiso/p120-catenin and TCF/beta-catenin complexes coordinately regulate canonical Wnt gene targets. | Park JI et al |
| 16469707 | 2006 | p120-catenin mediates inflammatory responses in the skin. | Perez-Moreno M et al |
| 12640114 | 2003 | p120 Catenin-associated Fer and Fyn tyrosine kinases regulate beta-catenin Tyr-142 phosphorylation and beta-catenin-alpha-catenin Interaction. | Piedra J et al |
| 11445535 | 2001 | The p120 catenin partner Kaiso is a DNA methylation-dependent transcriptional repressor. | Prokhortchouk A et al |
| 2469003 | 1989 | Transformation-specific tyrosine phosphorylation of a novel cellular protein in chicken cells expressing oncogenic variants of the avian cellular src gene. | Reynolds AB et al |
| 19525934 | 2009 | Essential role for eIF4GI overexpression in the pathogenesis of inflammatory breast cancer. | Silvera D et al |
| 19015320 | 2008 | p120 catenin induces opposing effects on tumor cell growth depending on E-cadherin expression. | Soto E et al |
| 15817151 | 2005 | The catenin p120ctn inhibits Kaiso-mediated transcriptional repression of the beta-catenin/TCF target gene matrilysin. | Spring CM et al |
| 20151151 | 2010 | Altered expression of p120catenin predicts poor outcome in invasive breast cancer. | Talvinen K et al |
| 12492499 | 2002 | Altered expression of the catenin p120 in human cancer: implications for tumor progression. | Thoreson MA et al |
| 16935280 | 2006 | p120 serine and threonine phosphorylation is controlled by multiple ligand-receptor pathways but not cadherin ligation. | Xia X et al |
| 12885254 | 2003 | Adhesion-associated and PKC-modulated changes in serine/threonine phosphorylation of p120-catenin. | Xia X et al |
| 14610056 | 2003 | Cellular levels of p120 catenin function as a set point for cadherin expression levels in microvascular endothelial cells. | Xiao K et al |
| 10753936 | 2000 | Receptor protein-tyrosine phosphatase RPTPmu binds to and dephosphorylates the catenin p120(ctn). | Zondag GC et al |
| 17030444 | 2007 | Diverse functions of p120ctn in tumors. | van Hengel J et al |
Other Information
Locus ID:
NCBI: 1500
MIM: 601045
HGNC: 2515
Ensembl: ENSG00000198561
Variants:
dbSNP: 1500
ClinVar: 1500
TCGA: ENSG00000198561
COSMIC: CTNND1
RNA/Proteins
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 38796558 | 2024 | CTNND1 is involved in germline predisposition to early-onset gastric cancer by affecting cell-to-cell interactions. | 0 |
| 38796558 | 2024 | CTNND1 is involved in germline predisposition to early-onset gastric cancer by affecting cell-to-cell interactions. | 0 |
| 37209565 | 2023 | The E3 ubiquitin ligase MIB1 suppresses breast cancer cell migration through regulating CTNND1 protein level. | 0 |
| 38134879 | 2023 | Tag, you're it: G-ICAT identifies p120-cysteine glutathionylation triggering E-cadherin destabilization. | 0 |
| 37209565 | 2023 | The E3 ubiquitin ligase MIB1 suppresses breast cancer cell migration through regulating CTNND1 protein level. | 0 |
| 38134879 | 2023 | Tag, you're it: G-ICAT identifies p120-cysteine glutathionylation triggering E-cadherin destabilization. | 0 |
| 34852711 | 2022 | Downregulated circular RNA hsa_circ_0005797 inhibits endometrial cancer by modulating microRNA-298/Catenin delta 1 signaling. | 7 |
| 35660718 | 2022 | Nuclear p120 catenin is a component of the perichromosomal layer and coordinates sister chromatid segregation during mitosis in lung cancer cells. | 0 |
| 35700046 | 2022 | CTNND1 variants cause familial exudative vitreoretinopathy through the Wnt/cadherin axis. | 14 |
| 35934314 | 2022 | p120 regulates E-cadherin expression in nasal epithelial cells in chronic rhinosinusitis. | 2 |
| 34852711 | 2022 | Downregulated circular RNA hsa_circ_0005797 inhibits endometrial cancer by modulating microRNA-298/Catenin delta 1 signaling. | 7 |
| 35660718 | 2022 | Nuclear p120 catenin is a component of the perichromosomal layer and coordinates sister chromatid segregation during mitosis in lung cancer cells. | 0 |
| 35700046 | 2022 | CTNND1 variants cause familial exudative vitreoretinopathy through the Wnt/cadherin axis. | 14 |
| 35934314 | 2022 | p120 regulates E-cadherin expression in nasal epithelial cells in chronic rhinosinusitis. | 2 |
| 33237836 | 2021 | APC regulation of ESRP1 and p120-catenin isoforms in colorectal cancer cells. | 12 |
Citation
Michael R Dohn ; Albert B Reynolds
CTNND1 (catenin (cadherin-associated protein), delta 1)
Atlas Genet Cytogenet Oncol Haematol. 2010-05-01
Online version: http://atlasgeneticsoncology.org/gene/40197/ctnnd1
