USP15 (ubiquitin specific peptidase 15)
2010-12-01 Monica Faronato , Sylvie Urbé , Judy M Coulson AffiliationPhysiology Department, School of Biomedical Sciences, Faculty of Health, Life Sciences, University of Liverpool, Crown Street, Liverpool, L69 3BX, UK
DNA/RNA
Note
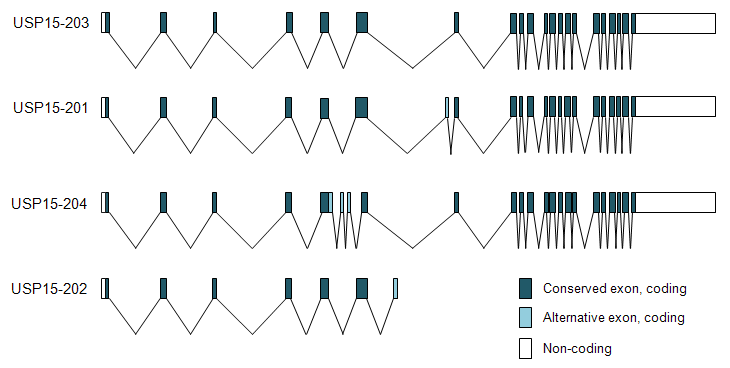
Description
Transcription
Summary table of USP15 transcripts.
| Name | Ensembl | Entrez | Aceview | Size (bp) | Exons |
| USP15-203 | ENST00000353364 | NM_006313.1 | Variant b | 4611 | 21 |
| USP15-201 | ENST00000280377 | Variant a | 4698 | 22 | |
| USP15-204 | ENST00000393654 | 4626 | 23 | ||
| USP15-202 | ENST00000312635 | Variant e | 717 | 7 |
Proteins
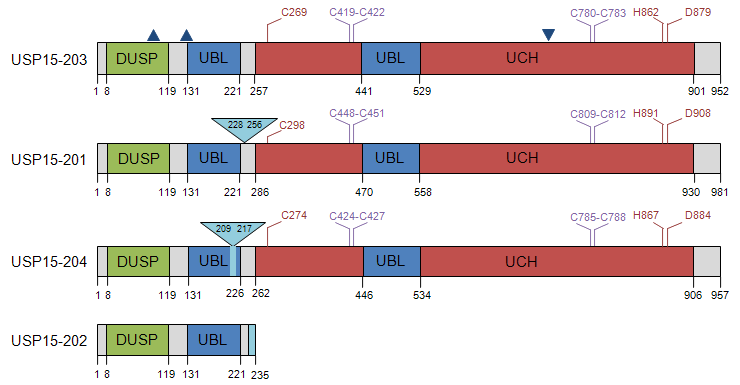
Description
| Name | Ensembl | Entrez | Size (aa) | MW (kDa) |
| USP15-203 | ENSP00000258123 | NP_006304.1 | 952 | 109 |
| USP15-201 | ENSP00000280377 | 981 | 112 | |
| USP15-204 | ENSP00000377264 | 957 | 109 | |
| USP15-202 | ENSP00000309240 | 235 | 40 |
All USP15 isoforms encompass a single N-terminal DUSP (domain in USPs) characterised by a novel tripod-like fold with a conserved hydrophobic surface patch that is predicted to participate in protein-protein interaction (de Jong et al., 2006). The ubiquitin carboxyl-terminal hydrolase (UCH) catalytic core of the USPs is typically around 350 amino acids, but consists of six conserved boxes, interspersed by insertion sites for additional sequence that can confer diversity (Ye et al., 2009). The major insertion in USP15 is between boxes 3/4, and embeds an ubiquitin-like fold (UBL) domain within the catalytic domain. UBLs are commonly found in the USPs and in certain other DUB families (Zhu et al., 2007; Komander et al., 2009a). They exhibit low sequence conservation, but have high structural similarity with ubiquitin and have been proposed to play important roles in regulating DUB catalytic function or interactions (Zhu et al., 2007; Ye et al., 2009). The intercalation of a UBL between boxes 3 and 4 of the catalytic domain increases the spacing between two sets of zinc-coordinating cysteine motifs, which form a functional zinc finger that is required for activity (Hetfeld et al., 2005). In the case of USP15, a second UBL is located directly adjacent to the DUSP (Zhu et al., 2007; Ye et al., 2009).
The four USP15 splice variants encode four distinct protein isoforms, which are illustrated in the diagram and summarised in the accompanying table. As a consequence of alternative splicing, isoform USP15-201 has a 29 amino acid insert within the unstructured region between the first UBL domain and the start of the UCH domain, whereas USP15-204 has a substitution of 3 amino acids for 8 residues within the first UBL. Otherwise the three isoforms that retain the catalytic domain are identical. They also retain predicted nuclear export signals (NES) (Soboleva et al., 2005) and, by homology with rat, a functional nuclear localisation signal (NLS) (Park et al., 2000).
Expression
Localisation
Function
USP15 has activity against both mono-ubiquitinated and poly-ubiquitinated substrates; the zinc-binding domain is necessary for USP15 to process poly-ubiquitin chains, but is not required for USP15 to remove ubiquitin from linear ubiquitin-GFP fusion proteins (Hetfeld et al., 2005). Although USP15 is relatively promiscuous in showing little specificity between K48- and K63-linked poly-ubiquitin chains, or between K63 and K11 di-ubiquitin linkages, it has limited activity against K11-linked poly-ubiquitin chains or linear ubiquitin (Komander et al., 2009b; Bremm et al., 2010).
A recent endeavour to map protein partners of the DUBs by mass spectroscopy reported that, in common with USP4 and USP39, USP15 interacts with several proteins involved in mRNA processing and so may play a role in ubiquitin-dependent control of splicing or mRNA decay (Sowa et al., 2009). In addition, various cancer-signalling pathways have been associated with USP15. For example, USP15 was one of twelve DUBs identified from an siRNA screen that impact on the hepatocyte growth factor (HGF)-dependent cell scattering response in non-small cell lung cancer and pancreatic cancer cells (Buus et al., 2009). A number of specific USP15 substrates have also been described, including the human papilloma virus (HPV) E6 oncoprotein (Vos et al., 2009), the RING-box protein Rbx1 (Hetfeld et al., 2005), the adenomatous polyposis coli (APC) tumour suppressor (Huang et al., 2009), and the NF-kB inhibitor IkBa (Schweitzer et al., 2007).
The latter three examples are all connected with the COP9-signalosome (CSN), a conserved multi-protein complex that regulates the cullin-RING ligase (CRL) superfamily of ubiquitin E3 ligases (Wei et al., 2008). CRLs have a core complex comprised of a cullin scaffold and the RING-box protein Rbx1 that recruit alternative adapter and substrate recognition proteins to form diverse E3 complexes with different substrate specificities. The primary function of the CSN is to remove the ubiquitin-like modifier Nedd8 from the cullin component. This both terminates E3 activity and is required for the reassembly of new CRLs (Wei et al., 2008). The CSN plays a role in many cancer-associated pathways including the cell cycle and DNA damage repair, and both CSN and CRL components may be dysregulated in tumours (Richardson and Zundel, 2005). Ubp12p, an S. pombe ortholog of human USP15, was shown through a systematic mass spectrometry screen to bind the CSN (Zhou et al., 2003). This targets Ubp12p to nuclear cullins, where it is proposed to protect against auto-ubiquitination and degradation of CRL components, in particular the substrate-specific adaptors. (Zhou et al., 2003; Wee et al., 2005).
Human USP15 also co-purifies with the CSN complex and was reported to stabilise the CRL core component Rbx1 (Hetfeld et al., 2005), thereby acting as a positive regulator of these E3 ligase complexes. In contrast, other studies suggest USP15 may directly oppose CRL E3 ligase activity by deubiquitinating specific substrates. For example, the CSN is involved in ubiquitin-dependent turnover of the IkBa inhibitor that retains NF-kB in the cytosol (Schweitzer et al., 2007). Phosphorylation of IkBa triggers CRL-mediated poly-ubiquitination of IkBa and subsequent proteasomal degradation, allowing NF-kB to enter the nucleus and activate transcription (Karin and Ben-Neriah, 2000). In response to TNFalpha, IkBa has been reported to interact with the CSN leading to its deubiquitination and stabilisation by CSN-associated USP15 (Schweitzer et al., 2007).
The adenomatous polyposis coli (APC) tumour suppressor and the beta-catenin oncogene are frequently mutated in cancers, particularly of the intestine, leading to constitutive wingless and Int-1 (Wnt) signalling (Clevers, 2006). The CSN is proposed to control the balance of beta-catenin and APC through formation of a regulatory super-complex. Deneddylation by the CSN promotes assembly of the beta-catenin destruction complex, whilst CSN-associated USP15 stabilises APC (Huang et al., 2009). The APC also plays a role in mitotic fidelity through interaction with the plus end-binding protein EB1 that controls microtubule growth and dynamics. In contrast to APC, EB1 is destabilised by USP15, suggesting that this is not a direct substrate, but rather that USP15 stabilises a CRL that accelerates ubiquitination and degradation of EB1 (Peth et al., 2007). It is interesting to speculate that such links with microtubule regulation may underpin recent reports that USP15 levels can influence the taxol sensitivity of cancer cells (Xu et al., 2009; Xie et al., 2010).
VCP/p97 is a large AAA+-type ATPase that acts as a chaperone in many cellular processes. Its basic function is to segregate ubiquitinated proteins from macromolecular complexes, and VCP plays an important role in recognizing and handling misfolded proteins, which are then either handed over for degradation or recycled. The CSN directly interacts with VCP and USP15 can process VCP-bound poly-ubiquitinated substrates, which accumulate following USP15 depletion (Cayli et al., 2009). VCP is implicated in human neurodegenerative disorders where it co-localises with poly-glutamine aggregates and is proposed to act as both an aggregate-formase and an unfoldase (Kakizuka, 2008). Another established VCP-associated cofactor, the DUB Ataxin-3, is subject to polyglutamine repeat expansion, which causes Machado-Joseph disease (Madsen et al., 2009). Although the mechanism is as yet unclear, USP15 was recently associated with this same disorder (Menzies et al., 2010).
Homology
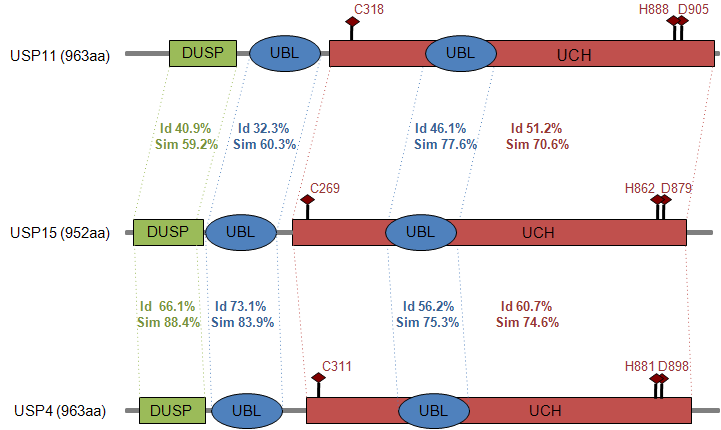
Mutations
Somatic
Implicated in
Germline mutations in APC lead to inherited colon cancer and sporadic tumours are associated with beta-catenin stabilisation. Huang et al. show a role for USP15 in stabilizing APC levels through the action of the CSN (Huang et al., 2009).
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 15571815 | 2004 | Mechanism and function of deubiquitinating enzymes. | Amerik AY et al |
| 12532266 | 2003 | Isolation and characterization of the mouse ubiquitin-specific protease Usp15. | Angelats C et al |
| 10444327 | 1999 | Identification, functional characterization, and chromosomal localization of USP15, a novel human ubiquitin-specific protease related to the UNP oncoprotein, and a systematic nomenclature for human ubiquitin-specific proteases. | Baker RT et al |
| 11571651 | 2001 | Association of UNP, a ubiquitin-specific protease, with the pocket proteins pRb, p107 and p130. | Blanchette P et al |
| 20622874 | 2010 | Lys11-linked ubiquitin chains adopt compact conformations and are preferentially hydrolyzed by the deubiquitinase Cezanne. | Bremm A et al |
| 19699092 | 2009 | Deubiquitinase activities required for hepatocyte growth factor-induced scattering of epithelial cells. | Buus R et al |
| 19826004 | 2009 | COP9 signalosome interacts ATP-dependently with p97/valosin-containing protein (VCP) and controls the ubiquitination status of proteins bound to p97/VCP. | Cayli S et al |
| 17081971 | 2006 | Wnt/beta-catenin signaling in development and disease. | Clevers H et al |
| 11571652 | 2001 | The de-ubiquitinating enzyme Unp interacts with the retinoblastoma protein. | DeSalle LM et al |
| 16005295 | 2005 | The zinc finger of the CSN-associated deubiquitinating enzyme USP15 is essential to rescue the E3 ligase Rbx1. | Hetfeld BK et al |
| 19576224 | 2009 | The COP9 signalosome mediates beta-catenin degradation by deneddylation and blocks adenomatous polyposis coli destruction via USP15. | Huang X et al |
| 18208395 | 2008 | Roles of VCP in human neurodegenerative disorders. | Kakizuka A et al |
| 10837071 | 2000 | Phosphorylation meets ubiquitination: the control of NF-[kappa]B activity. | Karin M et al |
| 19373254 | 2009 | Molecular discrimination of structurally equivalent Lys 63-linked and linear polyubiquitin chains. | Komander D et al |
| 18408009 | 2008 | USP11 stabilizes HPV-16E7 and further modulates the E7 biological activity. | Lin CH et al |
| 19497384 | 2009 | New ATPase regulators--p97 goes to the PUB. | Madsen L et al |
| 20007218 | 2010 | Autophagy induction reduces mutant ataxin-3 levels and toxicity in a mouse model of spinocerebellar ataxia type 3. | Menzies FM et al |
| 10880343 | 2000 | Tissue-specificity, functional characterization and subcellular localization of a rat ubiquitin-specific processing protease, UBP109, whose mRNA expression is developmentally regulated. | Park KC et al |
| 17350042 | 2007 | Ubiquitin-dependent proteolysis of the microtubule end-binding protein 1, EB1, is controlled by the COP9 signalosome: possible consequences for microtubule filament stability. | Peth A et al |
| 15571809 | 2004 | Ubiquitin: structures, functions, mechanisms. | Pickart CM et al |
| 16380502 | 2005 | The emerging role of the COP9 signalosome in cancer. | Richardson KS et al |
| 20073038 | 2010 | Emerging roles of deubiquitinases in cancer-associated pathways. | Sacco JJ et al |
| 17318178 | 2007 | CSN controls NF-kappaB by deubiquitinylation of IkappaBalpha. | Schweitzer K et al |
| 15494318 | 2005 | Nuclear-cytoplasmic shuttling of the oncogenic mouse UNP/USP4 deubiquitylating enzyme. | Soboleva TA et al |
| 19615732 | 2009 | Defining the human deubiquitinating enzyme interaction landscape. | Sowa ME et al |
| 18687060 | 2008 | Protein partners of deubiquitinating enzymes. | Ventii KH et al |
| 19553310 | 2009 | The ubiquitin-specific peptidase USP15 regulates human papillomavirus type 16 E6 protein stability. | Vos RM et al |
| 15793566 | 2005 | CSN facilitates Cullin-RING ubiquitin ligase function by counteracting autocatalytic adapter instability. | Wee S et al |
| 18926707 | 2008 | The COP9 signalosome: more than a protease. | Wei N et al |
| 20651371 | 2010 | CXCR4, a potential predictive marker for docetaxel sensitivity in gastric cancer. | Xie L et al |
| 19665996 | 2009 | USP15 plays an essential role for caspase-3 activation during Paclitaxel-induced apoptosis. | Xu M et al |
| 19734957 | 2009 | Dissection of USP catalytic domains reveals five common insertion points. | Ye Y et al |
| 12718879 | 2003 | Fission yeast COP9/signalosome suppresses cullin activity through recruitment of the deubiquitylating enzyme Ubp12p. | Zhou C et al |
| 17597129 | 2007 | High incidence of ubiquitin-like domains in human ubiquitin-specific proteases. | Zhu X et al |
| 16298993 | 2006 | Solution structure of the human ubiquitin-specific protease 15 DUSP domain. | de Jong RN et al |
Other Information
Locus ID:
NCBI: 9958
MIM: 604731
HGNC: 12613
Ensembl: ENSG00000135655
Variants:
dbSNP: 9958
ClinVar: 9958
TCGA: ENSG00000135655
COSMIC: USP15
RNA/Proteins
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 38656882 | 2024 | USP15 facilitates the progression of bladder cancer by amplifying the activation of the NF-κB signaling pathway. | 0 |
| 38902237 | 2024 | In vivo CRISPR screens reveal SCAF1 and USP15 as drivers of pancreatic cancer. | 0 |
| 38656882 | 2024 | USP15 facilitates the progression of bladder cancer by amplifying the activation of the NF-κB signaling pathway. | 0 |
| 38902237 | 2024 | In vivo CRISPR screens reveal SCAF1 and USP15 as drivers of pancreatic cancer. | 0 |
| 36640492 | 2023 | Over-expression of USP15/MMP3 predict poor prognosis and promote growth, migration in non-small cell lung cancer cells. | 4 |
| 37012401 | 2023 | Loss of the receptors ER, PR and HER2 promotes USP15-dependent stabilization of PARP1 in triple-negative breast cancer. | 3 |
| 37373211 | 2023 | USP15-USP7 Axis and UBE2T Differential Expression May Predict Pathogenesis and Poor Prognosis in De Novo Myelodysplastic Neoplasm. | 1 |
| 37743483 | 2023 | ACVRL1 drives resistance to multitarget tyrosine kinase inhibitors in colorectal cancer by promoting USP15-mediated GPX2 stabilization. | 0 |
| 36640492 | 2023 | Over-expression of USP15/MMP3 predict poor prognosis and promote growth, migration in non-small cell lung cancer cells. | 4 |
| 37012401 | 2023 | Loss of the receptors ER, PR and HER2 promotes USP15-dependent stabilization of PARP1 in triple-negative breast cancer. | 3 |
| 37373211 | 2023 | USP15-USP7 Axis and UBE2T Differential Expression May Predict Pathogenesis and Poor Prognosis in De Novo Myelodysplastic Neoplasm. | 1 |
| 37743483 | 2023 | ACVRL1 drives resistance to multitarget tyrosine kinase inhibitors in colorectal cancer by promoting USP15-mediated GPX2 stabilization. | 0 |
| 34465865 | 2022 | The deubiquitinase USP15 modulates cellular redox and is a therapeutic target in acute myeloid leukemia. | 12 |
| 35422093 | 2022 | USP15 negatively regulates lung cancer progression through the TRAF6-BECN1 signaling axis for autophagy induction. | 18 |
| 34465865 | 2022 | The deubiquitinase USP15 modulates cellular redox and is a therapeutic target in acute myeloid leukemia. | 12 |
Citation
Monica Faronato ; Sylvie Urbé ; Judy M Coulson
USP15 (ubiquitin specific peptidase 15)
Atlas Genet Cytogenet Oncol Haematol. 2010-12-01
Online version: http://atlasgeneticsoncology.org/gene/44585/usp15
