dic(17;20)(p11.2;q11.2)
2008-09-01 Aurelia M Meloni-Ehrig Affiliation1.Cytogenetics/Oncology, Quest Diagnostics Nichols Institute, 14225 Newbrook Drive, Chantilly, VA 20151, USA
Clinics and Pathology
Disease
Phenotype stem cell origin
8 cases reported: 4 de novo AML cases (including one AML-M2 and one erythroleukemia ), 3 de novo MDS cases (including one refractory anemia ), one t-MDS, and one t-MDS in transformation to AMMoL.
Epidemiology
Epidemiology of the 8 patients reported to date, 7 were male and one was female, aged 47 to 87 yrs.
Prognosis
Poor; majority of patients died between 2 and 8 months post diagnosis.
Cytogenetics
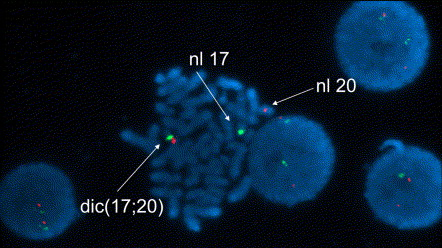
FISH image of a metaphase showing a normal copy of chromosomes 17 and 20 as well as the dic(17;20). The metaphase appears stained in blue (DAPI counterstain). Red signal, chromosome 20 centromere; green signal: chromosome 17 centromere. The dic(17;20) shows both centromeres.
Additional anomalies
Sole anomaly in one case; remaining cases with additional abnormalities; association with -5/del(5q), -7/del(7q), and/or +8 is frequent.
Genes Involved and Proteins
Note
dic(17;20) leads to loss of 17p (TP53 gene). Because of this, patients with this abnormality may have a prognostic outcome similar to the patients with "17p- syndrome". Dicentric (17;20) also leads to loss of 20q [various genes involved: topoisomerase 1 ( TOP1 ), phospholipase C ( PLC1 ), hepatocyte factor nuclear 4 ( HNF4 ), adenosine deaminase ( ADA ); KRML transcriptional regulator].
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 9130624 | 1997 | The 17p-syndrome: a distinct myelodysplastic syndrome entity? | Jary L et al |
| 12419582 | 2002 | A new nonrandom unbalanced t(17;20) in myeloid malignancies. | Patsouris C et al |
| 2120890 | 1990 | Granulocyte maturation and the chromosome deletion 17p- in primary myelodysplastic syndrome. | Pedersen B et al |
| 10748299 | 2000 | 17p- syndrome arising from a novel dicentric translocation in a patient with acute myeloid leukemia. | Watson N et al |
Summary
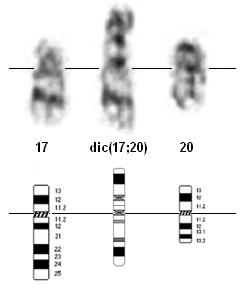
Dicentric (17;20)(p11.2;q11.2) partial karyotype and ideogram.
Citation
Aurelia M Meloni-Ehrig
dic(17;20)(p11.2;q11.2)
Atlas Genet Cytogenet Oncol Haematol. 2008-09-01
Online version: http://atlasgeneticsoncology.org/haematological/1485/dic(17;20)(p11-2;q11-2)
