RHOBTB2 (Rho-related BTB domain containing 2)
2016-02-01 Kristina Schenková , Shuo Cai , Francisco Rivero AffiliationIdentity
Abstract
RhoBTB2 is one of the three members of the RhoBTB family. All RhoBTB proteins are characterized by a GTPase domain followed by a proline-rich region, a tandem of two BTB domains and a C-terminal putative RING finger domain. RHOBTB2 is a putative tumour suppressor gene. Expression of RHOBTB2 has been found decreased in breast, lung and bladder tumors, head and neck squamous cell carcinoma and osteosarcoma. Decreased expression is often the result of promoter methylation. Mutations are uncommon. The role of RhoBTB2 in tumorigenesis is unknown but may be related to its function as a component of Cullin 3-dependent ubiquitin ligase complexes regulating the cell cycle and apoptosis.
DNA/RNA
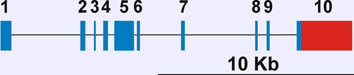
Description
Transcription
Proteins
Note
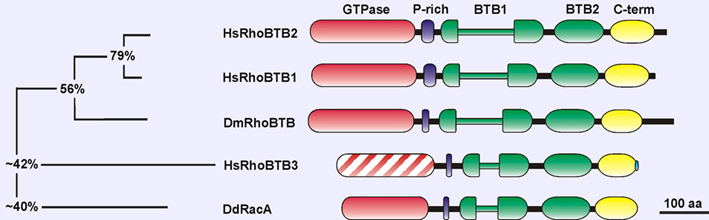
Description
The GTPase domain is Rho-related and contains a Rho insert that is longer than usual, two insertions and one deletion, as well as a few deviations from the GTPase consensus of most Rho GTPases. Chang et al. (2006) reported the inability of this domain to bind GTP. However the construct used in that study lacked one of the GTPase motifs, G5. Subsequent work with the full-length protein and a more complete GTPase domain showed that this domain is actually able to bind GTP (Manjarrez et al., 2014).
The proline-rich region links the GTPase to the first BTB domain. This region could act as a SH3 domain-binding site.
The BTB domain (broad complex, tramtract and bric-a-brac) is an evolutionary conserved protein-protein interaction domain that participates in homomeric and heteromeric associations with other BTB domains. The BTB domain was also identified as a component of multimeric cullin3-dependent ubiquitin ligase complexes. The first BTB domain is bipartite, being interrupted by an insertion of unknown function. The BTB domains of RhoBTB allow the formation of homodimers and of heterodimers with other proteins of the RhoBTB family (Berthold et al., 2008).
The C-terminus is a region conserved in all members of the RhoBTB subfamily. It predictably folds as 4 consecutive alpha-helices and one beta-strand and may constitute a RING finger domain (Manjarrez et al., 2014). Many RING finger domains function as ubiquitin ligases. RhoBTB2 does not bear a CAAX motif that is typical for classical Rho GTPases and serves for localization of the protein to membranes.
Expression
RhoBTB2 levels increase upon initiation of prophase and decrease at telophase. RhoBTB2 levels also increase during drug-induced apoptosis. Both effects depend on the E2F1 transcription factor (Freeman et al., 2007).
RHOBTB2 is a TP53 candidate target gene (Garritano et al., 2013).
Expression of RHOBTB2 has been found decreased in breast, lung, bladder and stomach cancers and in osteosarcomas (Hamaguchi et al., 2002; Knowles et al., 2005; Cho et al., 2008; Shi et al., 2008; Dong et al., 2012; Han et al., 2013; Jin et al., 2013) as well as in cell lines derived from breast, lung and bladder tumors and head and neck squamous cell carcinomas (Hamaguchi et al., 2002; Knowles et al., 2005; McKinnon et al., 2008) Loss of RHOBTB2 expression has been found to correlate with promoter methylation in breast and bladder cancers (Shi et al., 2008; Mehri Hajikhan et al., 2012; Han et al., 2013).
Localisation
Function
Following functions have been proposed for RhoBTB2:
1. RhoBTB2 as adaptor of cullin3-dependent ubiquitin ligases. The first BTB domain binds to the N-terminal region of Cullin 3, but not other cullins. RhoBTB2 is itself a substrate for the Cullin 3-based ubiquitin ligase complex (Wilkins et al., 2004). RhoBTB proteins appear to exist in an inactive state through an intramolecular interaction of the BTB domain region with the GTPase domain (Berthold et al., 2008). This model has been refined recently to show that the HSP90AA1 (Hsp90) chaperone machinery unlocks RhoBTB, enabling GTP binding and interaction with Cullin 3 and the COPS8 (COP9) signalosome. COP9 deneddylates Cullin 3 and stabilises the complex (Manjarrez et al. 2014).
RhoBTB2, like RhoBTB1 and RhoBTB3, interacts with LLRC41 (leucine rich repeat containing 41, MUF1). MUF1 is a nuclear protein and carries a BC-box that functions as a linker in multicomponent Cullin 5-dependent ubiquitin ligase complexes (Schenková et al., 2012). MUF1 may be a substrate for RhoBTB-Cullin 3 ubiquitin ligase complexes. The function of MUF1 is unknown, but it is suspected to be involved in the DNA damage response.
2. RhoBTB2, cell growth and apoptosis. Overexpression of RhoBTB2 in the breast cancer cell line T-47D (a cell line that lacks RHOBTB2 transcripts) effectively suppressed cell growth in vitro (Hamaguchi et al., 2002). Overexpression of RhoBTB2 in osteosarcoma cells significantly arrested cells at G1 and resulted in apoptosis (Jin et al., 2013). In the thyroid carcinoma cell line SW579 treatment with recombinant RhoBTB2 for 24 hours inhibited proliferation and provoked an increase of the apoptotic ratio through the mitochondrial apoptotic signalling pathway (Wang et al., 2015), but its not clear how the exogenously added protein exerts those actions.
It was shown that overexpression of RhoBTB2 leads to a short-term increase in cell cycle progression and proliferation, but long-term expression has a negative effect on proliferation (Freeman et al., 2007). The growth arrest effect has been explained by the downregulation of CCND1 (cyclin D1). Cyclin D1 is upstream of cyclin E, and the overexpression of any of both prevented the growth arrest effect of RhoBTB2 (Yoshihara et al., 2007). The effect on cyclin D1 is probably post-transcriptional, but only partially dependent on proteasomal degradation (Collado et al., 2007). RHOBTB2 has been identified as a target of the E2F1 transcription factor. RhoBTB2 levels also increase during drug-induced apoptosis in an E2F1-dependent manner, and the downregulation of RHOBTB2 delays the onset of apoptosis (Freeman et al., 2007).
3. RhoBTB2 and chemokine expression. Downregulation of RhoBTB2 by RNA interference in primary lung epithelial cells causes a decrease in CXCL14 mRNA expression. The same effect was observed in keratinocytes and is apparently independent of Cullin 3-mediated protein degradation (McKinnon et al., 2008).
4. RhoBTB2 and vesicle transport. Knockdown of endogenous RhoBTB2 hindered the ER to Golgi apparatus transport of a VSVG-GFP reporter and resulted in the altered distribution of the fusion protein. Ectopic RhoBTB2 distributes in a vesicular pattern occasionally adjacent to microtubules and an intact microtubule network seems to be required for the mobility of RhoBTB2 (Chang el al., 2006).
5. RhoBTB2 and the actin filament system. RhoBTB2 displays only a moderate influence on the morphology and actin organisation of porcine aortic endothelial cells upon ectopic expression. It does not interact with the GTPase-binding domain of WASP, PAK1 or RTKN (Rhotekin), which are well-known effectors of many typical Rho GTPases (Aspenström et al., 2004).
Homology
Mutations
Note
| Mutation | Effect | Tumour |
| Prom -238G>A* | Altered expression? | Breast |
| Prom -121C>T | Altered expression? | Breast |
| 5UTR +48G>A | Altered expression? | Breast |
| E5 C>T R275W | Unknown effect | Stomach |
| E5 T>G Y284D | Abolished binding to Cullin 3 | Lung (cell line) |
| E5 G>A D299N | No growth inhibition when re-expressed | Breast |
| E5 G>C E349D | Unknown effect | Bladder |
| E5 A>C D368A | Unknown effect | Breast (cell line) |
| E7 G>A G561S* | Unknown effect | Bladder |
| E9 C>A P647T | Unknown effect | Breast |
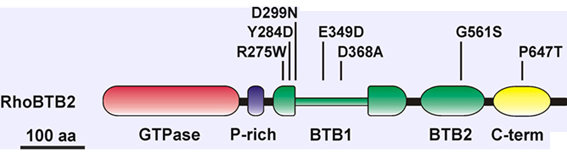
Implicated in
A more extensive mutation analysis of 100 sporadic breast cancers revealed some polymorphisms as well as two somatic mutations in the promoter (-238G>A, -121C>T) and 5UTR (+48G>A) of RHOBTB2. The analysis of 17 CC: TXT: familial breast tumours ID: 10062> negative for BRCA1/BRCA2mutations failed to reveal additional mutations in the coding region of RHOBTB2 (Ohadi et al., 2007).
No mutations were found in the promoter or exon 7 in 32 breast cancers of a Han Chinese population. This study revealed an intronic polymorphism common in this population; the variant IVS7 + 53C >G correlated with HER2 and p53 expression but not with age, tumor stage or estrogen or progesterone receptor expression (Fu et al., 2014.
Using semi-quantitative PCR RHOBTB2 mRNA was found absent (56% of 87 breast tumors vs. 9% of normal tissue) or significantly reduced. RHOBTB2 Promoter methylation was detected in over 33% of a large collection of breast tumor samples and was rare in normal tissue. Loss of RHOBTB2 expression correlated with promoter methylation. Moreover, RHOBTB2 promoter methylation associated with more advance tumor stages, p53 mutation and HER2-positive status (Han et al., 2013). A similar correlation between promoter methylation and RHOBTB2 downregulation has been reported in a sample of 50 paired breast cancer and normal tissues. In this study promoter methylation was found more frequently associated to progesterone receptor negative tumors (Tang et al., 2014). Significantly more frequent promoter methylation was also found in a collection of 50 breast tumor (34%) and blood samples (46%) compared to normal samples (20% or less) in another study (Mehri Hjikhan et al., 2012).
A frequent pattern of loss of RHOBTB2, CDH1 and TP53 together with gain of COX2 and MYC was found in a single cell in situ hybridization analysis of 13 samples simultaneously carrying ductal carcinoma in situ (DCIS) and invasive ductal carcinoma (IDC) of the breast. DCIS is the precursor lesion for invasive breast cancer. The study aimed at understanding the dynamics of genomic alterations in the progression from DCIS to IDC and included several oncogenes and tumor suppressor genes (Heselmayer-Haddad et al., 2012).
The mutations found in the promoter and 5UTR of RHOBTB2 in some breast tumors might affect regulation of gene expression (Ohadi et al., 2007). Lack of RHOBTB2 expression in T-47D is apparently due to RHOBTB2 promoter methylation (Han et al., 2013). Methylation of RHOBTB2 and other genes in peripheral blood is a potential epigenetic marker for predicting the risk of breast cancer development (Khakpour et al., 2015).
Ectopic expression of RHOBTB2 in two human metastatic breast cancer cell lines, MDA-MB-231 and MDA-MB-435, inhibits cell migration and invasiveness through a mechanism that involves upregulation of the metastasis suppressor BRMS1 and decreased phosphorylation of EZR (ezrin) and Akt2 (Ling et al., 2010). Ezrin is a cytoskeleton and signaling molecule that regulates cell adhesion, migration and invasion. Akt2 is a kinase involved in invasiveness of breast cancer cells and is able to phosphorylate ezrin (Freeman and Cress 2010)
In an immunohistochemistry study of 172 tissue samples from different subtypes of lung adenocarcinomas frequent (70% of tumors) downregulation of RHOBTB2 was found that correlated with the degree of invasiveness (Dong et al, 2012).
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 24485767 | 2014 | The mutation of DBC2 in breast cancer patients from the Han ethnic group in Eastern China. | Fu G et al |
| 24356943 | 2014 | RhoBTB2 gene in breast cancer is silenced by promoter methylation. | Tang W et al |
| 14521508 | 2004 | Rho GTPases have diverse effects on the organization of the actin filament system. | Aspenström P et al |
| 18298893 | 2008 | Rho GTPases of the RhoBTB subfamily and tumorigenesis. | Berthold J et al |
| 17023000 | 2006 | DBC2 is essential for transporting vesicular stomatitis virus glycoprotein. | Chang FK et al |
| 17906984 | 2008 | Genetic analysis of the DBC2 gene in gastric cancer. | Cho YG et al |
| 17617377 | 2007 | DBC2 resistance is achieved by enhancing 26S proteasome-mediated protein degradation. | Collado D et al |
| 17100600 | 2006 | Distinct and Overlapping Roles for E2F Family Members in Transcription, Proliferation and Apoptosis. | DeGregori J et al |
| 22901165 | 2012 | Loss of DBC2 expression is an early and progressive event in the development of lung adenocarcinoma. | Dong W et al |
| 20980811 | 2010 | RhoBTB2 (DBC2) comes of age as a multifunctional tumor suppressor. | Freeman SN et al |
| 23817466 | 2013 | More targets, more pathways and more clues for mutant p53. | Garritano S et al |
| 23626933 | 2012 | Evaluation of Methylation Status in the 5'UTR Promoter Region of the DBC2 Gene as a Biomarker in Sporadic Breast Cancer. | Hajikhan Mirzaei M et al |
| 12370419 | 2002 | DBC2, a candidate for a tumor suppressor gene involved in breast cancer. | Hamaguchi M et al |
| 23546941 | 2013 | Decreased expression of the DBC2 gene and its clinicopathological significance in breast cancer: correlation with aberrant DNA methylation. | Han L et al |
| 23062488 | 2012 | Single-cell genetic analysis of ductal carcinoma in situ and invasive breast cancer reveals enormous tumor heterogeneity yet conserved genomic imbalances and gain of MYC during progression. | Heselmeyer-Haddad K et al |
| 23777252 | 2013 | Downregulated RhoBTB2 expression contributes to poor outcome in osteosarcoma patients. | Jin Z et al |
| 26076810 | 2015 | DNA methylation as a promising landscape: A simple blood test for breast cancer prediction. | Khakpour G et al |
| 15922864 | 2005 | Mutation analysis of the 8p candidate tumour suppressor genes DBC2 (RHOBTB2) and LZTS1 in bladder cancer. | Knowles MA et al |
| 20930524 | 2010 | Ectopic expression of RhoBTB2 inhibits migration and invasion of human breast cancer cells. | Ling LJ et al |
| 24608665 | 2014 | Hsp90-dependent assembly of the DBC2/RhoBTB2-Cullin3 E3-ligase complex. | Manjarrez JR et al |
| 21801820 | 2011 | RhoBTB2 (DBC2) functions as tumor suppressor via inhibiting proliferation, preventing colony formation and inducing apoptosis in breast cancer cells. | Mao H et al |
| 18762809 | 2008 | The atypical Rho GTPase RhoBTB2 is required for expression of the chemokine CXCL14 in normal and cancerous epithelial cells. | McKinnon CM et al |
| 10048485 | 1998 | Prediction of the coding sequences of unidentified human genes. XII. The complete sequences of 100 new cDNA clones from brain which code for large proteins in vitro. | Nagase T et al |
| 17653899 | 2007 | Mutation analysis of the DBC2 gene in sporadic and familial breast cancer. | Ohadi M et al |
| 12426103 | 2002 | Genomic organization and expression profile of the small GTPases of the RhoBTB family in human and mouse. | Ramos S et al |
| 11222756 | 2001 | The Dictyostelium discoideum family of Rho-related proteins. | Rivero F et al |
| 22709582 | 2012 | MUF1/leucine-rich repeat containing 41 (LRRC41), a substrate of RhoBTB-dependent cullin 3 ubiquitin ligase complexes, is a predominantly nuclear dimeric protein. | Schenková K et al |
| 18640857 | 2008 | DBC2 gene is silenced by promoter methylation in bladder cancer. | Shi Y et al |
| 15663929 | 2005 | DBC2 significantly influences cell-cycle, apoptosis, cytoskeleton and membrane-trafficking pathways. | Siripurapu V et al |
| 15567721 | 2004 | High expression during neurogenesis but not mammogenesis of a murine homologue of the Deleted in Breast Cancer2/Rhobtb2 tumor suppressor. | St-Pierre B et al |
| 15107402 | 2004 | RhoBTB2 is a substrate of the mammalian Cul3 ubiquitin ligase complex. | Wilkins A et al |
| 17517369 | 2007 | Cyclin D1 down-regulation is essential for DBC2's tumor suppressor function. | Yoshihara T et al |
Other Information
Locus ID:
NCBI: 23221
MIM: 607352
HGNC: 18756
Ensembl: ENSG00000008853
Variants:
dbSNP: 23221
ClinVar: 23221
TCGA: ENSG00000008853
COSMIC: RHOBTB2
RNA/Proteins
| Gene ID | Transcript ID | Uniprot |
|---|---|---|
| ENSG00000008853 | ENST00000251822 | Q9BYZ6 |
| ENSG00000008853 | ENST00000519685 | Q9BYZ6 |
| ENSG00000008853 | ENST00000522948 | Q9BYZ6 |
| ENSG00000008853 | ENST00000524077 | E5RI44 |
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 37165955 | 2023 | Genotype-phenotype correlations in RHOBTB2-associated neurodevelopmental disorders. | 0 |
| 37165955 | 2023 | Genotype-phenotype correlations in RHOBTB2-associated neurodevelopmental disorders. | 0 |
| 36195378 | 2022 | Mild head trauma: Acute encephalopathy trigger in children with RHOBTB2 de novo mutation. | 0 |
| 36195378 | 2022 | Mild head trauma: Acute encephalopathy trigger in children with RHOBTB2 de novo mutation. | 0 |
| 33504645 | 2021 | RHOBTB2 Mutations Expand the Phenotypic Spectrum of Alternating Hemiplegia of Childhood. | 8 |
| 34074803 | 2021 | Identification of RHOBTB2 aberration as an independent prognostic indicator in acute myeloid leukemia. | 6 |
| 33504645 | 2021 | RHOBTB2 Mutations Expand the Phenotypic Spectrum of Alternating Hemiplegia of Childhood. | 8 |
| 34074803 | 2021 | Identification of RHOBTB2 aberration as an independent prognostic indicator in acute myeloid leukemia. | 6 |
| 29276004 | 2018 | Missense Variants in RHOBTB2 Cause a Developmental and Epileptic Encephalopathy in Humans, and Altered Levels Cause Neurological Defects in Drosophila. | 32 |
| 29276004 | 2018 | Missense Variants in RHOBTB2 Cause a Developmental and Epileptic Encephalopathy in Humans, and Altered Levels Cause Neurological Defects in Drosophila. | 32 |
| 27941885 | 2017 | DBC2/RhoBTB2 functions as a tumor suppressor protein via Musashi-2 ubiquitination in breast cancer. | 23 |
| 27941885 | 2017 | DBC2/RhoBTB2 functions as a tumor suppressor protein via Musashi-2 ubiquitination in breast cancer. | 23 |
| 24356943 | 2014 | RhoBTB2 gene in breast cancer is silenced by promoter methylation. | 7 |
| 24485767 | 2014 | The mutation of DBC2 in breast cancer patients from the Han ethnic group in Eastern China. | 0 |
| 24608665 | 2014 | Hsp90-dependent assembly of the DBC2/RhoBTB2-Cullin3 E3-ligase complex. | 12 |
Citation
Kristina Schenková ; Shuo Cai ; Francisco Rivero
RHOBTB2 (Rho-related BTB domain containing 2)
Atlas Genet Cytogenet Oncol Haematol. 2016-02-01
Online version: http://atlasgeneticsoncology.org/gene/42109/rhobtb2
Historical Card
2008-12-01 RHOBTB2 (Rho-related BTB domain containing 2) by Kristina Schenková,Francisco Rivero Affiliation
