Bloom syndrome
2005-02-01 Mounira Amor-Guéret AffiliationInstitut Curie - Section de Recherche, UMR 2027 CNRS, Bâtiment 110, Centre Universitaire, F-91405 Orsay Cedex, France
Identity
Name
Bloom syndrome
Inheritance
autosomal recessive; frequency is about 2\/105 newborns in Ashkenazi Jews and in the Japanese (founder effect: affected persons descent from a common ancestor); much rarer otherwise
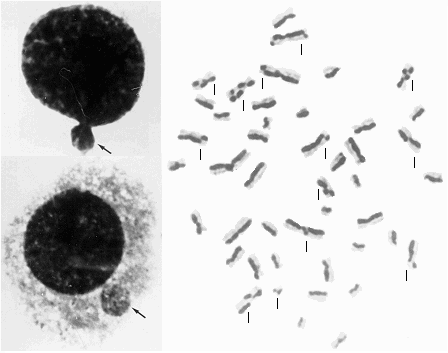
micronuclei (left); sister chromatid exchange (right) in a normal subject (herein: 19 SCE, instead of the hundred found in Bloom, see below) - Editor
Omim
210900
Mesh
D001816
Orphanet
125 Bloom syndrome
Umls
C0005859
Clinics
Note
168 cases have been registered in the Blooms syndrome Registry by James German; BS patients are predisposed to all types of cancer observed in the general population; thus, BS is a model of initiation and promotion of cancer, and highligths internal causes\/processes of cancers
Phenotype and clinics
- phenotypic spectrum variable.
- growth : dwarfism: intrauterine growth retardation; birth weight: below 2.3 kg; mean length: 44 cm; adult length < 145 cm.
- skin: hyperpigmented (café au lait) spots; hypopigmented areas; sun sensitive telangiectatic erythema; in butterfly configuration across the face: resembles lupus erythematous
- head: microcephaly; dolichocephaly; narrow face; prominent nose and\/or ears; characteristic high-pitched voice
- normal intelligence
- immune deficiency --> frequent infections (may be life-threatening)
- other: myocardopathy; hypogonadism in male patients; hypertriglyceridemia
- growth : dwarfism: intrauterine growth retardation; birth weight: below 2.3 kg; mean length: 44 cm; adult length < 145 cm.
- skin: hyperpigmented (café au lait) spots; hypopigmented areas; sun sensitive telangiectatic erythema; in butterfly configuration across the face: resembles lupus erythematous
- head: microcephaly; dolichocephaly; narrow face; prominent nose and\/or ears; characteristic high-pitched voice
- normal intelligence
- immune deficiency --> frequent infections (may be life-threatening)
- other: myocardopathy; hypogonadism in male patients; hypertriglyceridemia
Neoplastic risk
Evolution
major medical complications apart from cancers are : chronic lung disease, and diabetes mellitus (in 10 %)
Prognosis
1\/3 of patients are dead at mean age 24 yrs (oldest died at 49 yrs, youngest died before 1 yr) and the mean age of the 2\/3 remaining alive patients is 22 yrs (range: 4-46 yrs)
Cytogenetics
Inborn condition

sister chromatid exchange in a normal subject (left) and in a Bloom syndrome patient (right) (from: Mounira Amor-Guéret)
Other Findings
Note
Genes involved and Proteins
Complementation groups
no complementation group
Description
1417 amino acids; contains one ATP binding site, one DEAH box, and two putative nuclear localization signals
Expression
accumulates to high levels in S phase of the cell cycle, persists in G2\/M and sharply declines in G1; hyperphoshorylated in mitosis
Localisation
nuclear (PML nuclear bodies and nucleolus)
Function
Homology
with the RecQ helicases
Germinal
five BLM mutations introducing amino acid substitutions and four BLM mutations introducing premature nonsense codons into the coding sequence have been described to date; one BLM mutation consisting in a 6 bp deletion accompanied by a 7 bp insertion at nucleic acid position 2281 is common in patients from Ashkenazi Jewish ancestry, leading to a truncated protein of 739 amino acids in length; two BLM mutations, 631delCAA and 1610insA were detected in japanese patients.
To be noted
Hgmd
135698 BLM
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 11960380 | 2002 | Bloom's syndrome protein response to ultraviolet-C radiation and hydroxyurea-mediated DNA synthesis inhibition. | Ababou M et al |
| 10862105 | 2000 | Identification of a novel BLM missense mutation (2706T>C) in a Moroccan patient with Bloom's syndrome. | Barakat A et al |
| 15604258 | 2004 | A major role for mitotic CDC2 kinase inactivation in the establishment of the mitotic DNA damage checkpoint. | Bayart E et al |
| 11309417 | 2001 | Regulation and localization of the Bloom syndrome protein in response to DNA damage. | Bischof O et al |
| 10851073 | 2000 | Cell cycle regulation of the endogenous wild type Bloom's syndrome DNA helicase. | Dutertre S et al |
| 8875252 | 1996 | Molecular genetics of Bloom's syndrome. | Ellis NA et al |
| 7585968 | 1995 | The Bloom's syndrome gene product is homologous to RecQ helicases. | Ellis NA et al |
| 9285778 | 1997 | Characterization of a new BLM mutation associated with a topoisomerase II alpha defect in a patient with Bloom's syndrome. | Foucault F et al |
| 11781842 | 2001 | The Bloom syndrome protein interacts and cooperates with p53 in regulation of transcription and cell growth control. | Garkavtsev IV et al |
| 11971187 | 2002 | Increased error-prone non homologous DNA end-joining--a proposed mechanism of chromosomal instability in Bloom's syndrome. | Gaymes TJ et al |
| 9062585 | 1997 | Bloom's syndrome. XX. The first 100 cancers. | German J et al |
| 10525530 | 1999 | PML is critical for ND10 formation and recruits the PML-interacting protein daxx to this nuclear structure when modified by SUMO-1. | Ishov AM et al |
| 15289897 | 2004 | Relatively common mutations of the Bloom syndrome gene in the Japanese population. | Kaneko H et al |
| 9388480 | 1997 | BLM (the causative gene of Bloom syndrome) protein translocation into the nucleus by a nuclear localization signal. | Kaneko H et al |
| 9388193 | 1997 | The Bloom's syndrome gene product is a 3'-5' DNA helicase. | Karow JK et al |
| 10823897 | 2000 | The Bloom's syndrome gene product promotes branch migration of holliday junctions. | Karow JK et al |
| 12019152 | 2002 | The BLM helicase is necessary for normal DNA double-strand break repair. | Langland G et al |
| 12724401 | 2003 | A multiprotein nuclear complex connects Fanconi anemia and Bloom syndrome. | Meetei AR et al |
| 11433031 | 2001 | The Bloom's and Werner's syndrome proteins are DNA structure-specific helicases. | Mohaghegh P et al |
| 14576316 | 2003 | Possible anti-recombinogenic role of Bloom's syndrome helicase in double-strand break processing. | Onclercq-Delic R et al |
| 12181313 | 2002 | Telomere-binding protein TRF2 binds to and stimulates the Werner and Bloom syndrome helicases. | Opresko PL et al |
| 11691925 | 2001 | Direct association of Bloom's syndrome gene product with the human mismatch repair protein MLH1. | Pedrazzi G et al |
| 14577841 | 2003 | Telomere and ribosomal DNA repeats are chromosomal targets of the bloom syndrome DNA helicase. | Schawalder J et al |
| 15364958 | 2004 | Functional interaction between BLM helicase and 53BP1 in a Chk1-mediated pathway during S-phase arrest. | Sengupta S et al |
| 15509577 | 2004 | Genetic interactions between BLM and DNA ligase IV in human cells. | So S et al |
| 12444098 | 2002 | The Bloom syndrome helicase BLM interacts with TRF2 in ALT cells and promotes telomeric DNA synthesis. | Stavropoulos DJ et al |
| 9765292 | 1998 | The Bloom's syndrome helicase unwinds G4 DNA. | Sun H et al |
| 15579905 | 2005 | Human Bloom protein stimulates flap endonuclease 1 activity by resolving DNA secondary structure. | Wang W et al |
| 11399766 | 2001 | Functional interaction of p53 and BLM DNA helicase in apoptosis. | Wang XW et al |
| 10783165 | 2000 | BASC, a super complex of BRCA1-associated proteins involved in the recognition and repair of aberrant DNA structures. | Wang Y et al |
| 11278509 | 2001 | Potential role for the BLM helicase in recombinational repair via a conserved interaction with RAD51. | Wu L et al |
| 12080066 | 2002 | The processing of Holliday junctions by BLM and WRN helicases is regulated by p53. | Yang Q et al |
| 10779560 | 2000 | Nuclear structure in normal and Bloom syndrome cells. | Yankiwski V et al |
| 11087418 | 2000 | Binding and melting of D-loops by the Bloom syndrome helicase. | van Brabant AJ et al |
| 11919194 | 2002 | Colocalization, physical, and functional interaction between Werner and Bloom syndrome proteins. | von Kobbe C et al |
External Links
Citation
Mounira Amor-Guéret
Bloom syndrome
Atlas Genet Cytogenet Oncol Haematol. 2005-02-01
Online version: http://atlasgeneticsoncology.org/cancer-prone-disease/10002/bloom-syndrome
Historical Card
2000-09-01 Bloom syndrome by Mounira Amor-Guéret Affiliation
Institut Curie - Section de Recherche, UMR 2027 CNRS, Bâtiment 110, Centre Universitaire, F-91405 Orsay Cedex, France
1998-02-01 Bloom syndrome by Jean-Loup Huret Affiliation
Genetics, Dept Medical Information, University of Poitiers, CHU Poitiers Hospital, F-86021 Poitiers, France
