CSNK1A1 (casein kinase 1, alpha 1)
2008-05-01 Max Doerfel , Otmar Huber AffiliationDepartment of Laboratory Medicine, Pathobiochemistry, Charite - Universitatsmedizin Berlin, Campus Benjamin Franklin, Hindenburgdamm 30, 12203 Berlin, Germany
Identity
HGNC
LOCATION
5q32
LOCUSID
ALIAS
CK1,CK1a,CKIa,HEL-S-77p,HLCDGP1,PRO2975
FUSION GENES
DNA/RNA
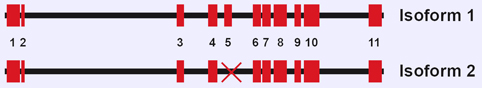
Diagram of hCK1 alpha gene. Exons for both isoforms are presented as red boxes, with exon mumbers at the bottom.
Description
According to Entrez-Gene, CNK1 alpha1 gene maps to NC_000005.8.
Transcription
In mammals, 7 distinct genes encoding CK1 isoforms (CK1 alpha, CK1 beta, CK1 gamma1, CK1 gamma2, CK1 gamma3, CK1 delta and CK1 epsilon) are expressed which differ mainly in length and primary structure of the C-terminal non-catalytic domain. Furthermore, CK1 alpha splice variants have been detected in many different organisms including vertebrates and mammals. Alternative splicing leads to the insertion of a long (L, 28 aa) or a short (S, 12 aa) insert into the catalytic C-terminal domain of CK1 alpha and generates different splicing products (CK1 alpha, CK1 alphaL, CK1 alphaS, CK2 alphaLS, alpha3) that differ in kinase activity, function and subcellular localization.
In humans at least two isoforms of CK1 alpha have been identified encoding a 3149 bp (isoform1/NM_001025105) or 3061 bp (isoform2/NM_001892) mRNA, respectively.
In humans at least two isoforms of CK1 alpha have been identified encoding a 3149 bp (isoform1/NM_001025105) or 3061 bp (isoform2/NM_001892) mRNA, respectively.
Proteins
Note
The hCK1 alpha isoforms are composed of 365 or 337 amino acids and have a calculated molecular weight of 41,9 or 38,9 kDa, respectively. Both isoforms contain a putative near-consensus SV40 T-antigen nuclear localization sequence.
Description
CK1 alpha is a second messenger-independent, monomeric, serine/threonine specific protein kinase that recognizes a canonical consensus sequence pS/pT-X1-2-S/T or (D/E)-X1-2-S/T. Additionally, a noncanonical sequence containing a SLS motif followed by a cluster of acidic residues C-terminal of the phosphoacceptor site is recognized by CK1.
Localisation
The protein kinase CK1 alpha is ubiquitously expressed in all tissues and detected in all cellular compartments.
Function
Mammalian CK1 isoforms and their splice variants are involved in diverse cellular processes including membrane trafficking, circadian rhythm, cell cycle progression, chromosome segregation, apoptosis and cellular proliferation and differentiation.
Implicated in
Entity name
Cell cycle control
Note
Entity name
Apoptosis
Note
Recently is has been shown, that CK1 is involved in negatively regulating apoptosis through phosphorylation of diverse cellular proteins including the p75 tumor necrosis factor, proteins of the death-inducing signaling complex (TRAIL induced apoptosis) or Bid (FAS-mediated apoptosis).
It is thought, that CK1 mediated phosphorylation at the level of death-inducing signaling complex (DISC) leads to resistance against caspase cleavage and thereby down regulation of TRAIL (tumor necrosis-factor-related apoptosis ligand) induced apoptosis.
Furthermore, there is evidence that CK1 alpha regulates Fas-mediated apoptosis through phosphorylation of the proapoptotic Bcl2 family member Bid, which prevents caspase 8-dependant cleavage of Bid and negatively influences Fas response.
Additionally, evidence has increased that CK1 alpha modulates RXR agonist mediated apoptosis through interaction and/or phosphorylation of RXR, which prevents cytochrome C realease from the mitochondria.
It is thought, that CK1 mediated phosphorylation at the level of death-inducing signaling complex (DISC) leads to resistance against caspase cleavage and thereby down regulation of TRAIL (tumor necrosis-factor-related apoptosis ligand) induced apoptosis.
Furthermore, there is evidence that CK1 alpha regulates Fas-mediated apoptosis through phosphorylation of the proapoptotic Bcl2 family member Bid, which prevents caspase 8-dependant cleavage of Bid and negatively influences Fas response.
Additionally, evidence has increased that CK1 alpha modulates RXR agonist mediated apoptosis through interaction and/or phosphorylation of RXR, which prevents cytochrome C realease from the mitochondria.
Entity name
Note
The Wnt pathway is a complex signaling cascade regulating cell proliferation and differentiation. During recent years, the significance of Wnt signaling in human cancer has been elucidated. Identification of numerous pathway components and mutations in the encoding genes finally result in stabilization and accumulation of beta-Catenin and enhanced transcription of TCF/LEF- beta-Catenin target genes.
CK1 alpha is part of the beta-Catenin destruction complex where it phosphorylates beta-Catenin at Serin 45, priming the subsequent phosphorylation of beta-Catenin by GSK3 beta. These phosphorylations mark beta-Catenin for proteasomal degradation. This is one of the central regulatory events controling the Wnt signaling-pathway.
Furthermore, CK1 has been shown to additionally regulate Wnt-signaling through phosphorylation of diverse cellular proteins including LEF-1 (lymphocyte enhancer factor-1) and beta-Catenin leading to the disruption of the LEF-1/ beta-Catenin transcription complex.
For additional information about Wnt-signaling in general, Wnt-signaling components and Wnt target genes, readers are referred to the Wnt-Homepage posted by the Nusse group.
CK1 alpha is part of the beta-Catenin destruction complex where it phosphorylates beta-Catenin at Serin 45, priming the subsequent phosphorylation of beta-Catenin by GSK3 beta. These phosphorylations mark beta-Catenin for proteasomal degradation. This is one of the central regulatory events controling the Wnt signaling-pathway.
Furthermore, CK1 has been shown to additionally regulate Wnt-signaling through phosphorylation of diverse cellular proteins including LEF-1 (lymphocyte enhancer factor-1) and beta-Catenin leading to the disruption of the LEF-1/ beta-Catenin transcription complex.
For additional information about Wnt-signaling in general, Wnt-signaling components and Wnt target genes, readers are referred to the Wnt-Homepage posted by the Nusse group.
Entity name
Neurodegenerative disorders
Note
Deregulation of CK1 expression and activity has been linked to various diseases including neurodegenerative disorders, especially in tauophathies like Alzheimers and Parkinsons disease.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 1409656 | 1992 | Cell cycle-dependent localization of casein kinase I to mitotic spindles. | Brockman JL et al |
| 10514399 | 1999 | A new molecular link between the fibrillar and granulovacuolar lesions of Alzheimer's disease. | Ghoshal N et al |
| 9884021 | 1998 | Casein kinase I: spatial organization and positioning of a multifunctional protein kinase family. | Gross SD et al |
| 9365278 | 1997 | A casein kinase I isoform is required for proper cell cycle progression in the fertilized mouse oocyte. | Gross SD et al |
| 15747065 | 2005 | A second protein kinase CK1-mediated step negatively regulates Wnt signalling by disrupting the lymphocyte enhancer factor-1/beta-catenin complex. | Hämmerlein A et al |
| 16557393 | 2006 | Casein kinase-1 isoforms differentially associate with neurofibrillary and granulovacuolar degeneration lesions. | Kannanayakal TJ et al |
| 16186692 | 2005 | The role of the casein kinase 1 (CK1) family in different signaling pathways linked to cancer development. | Knippschild U et al |
| 12925738 | 2003 | A noncanonical sequence phosphorylated by casein kinase 1 in beta-catenin may play a role in casein kinase 1 targeting of important signaling proteins. | Marin O et al |
| 12176352 | 2002 | Casein kinase 1: a Wnt'er of disconnect. | Polakis P et al |
| 16556058 | 2005 | The role of Wnt signaling in cancer and stem cells. | Reguart N et al |
Other Information
Locus ID:
NCBI: 1452
MIM: 600505
HGNC: 2451
Ensembl: ENSG00000113712
Variants:
dbSNP: 1452
ClinVar: 1452
TCGA: ENSG00000113712
COSMIC: CSNK1A1
RNA/Proteins
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 37723657 | 2024 | CSNK1A1/CK1α suppresses autoimmunity by restraining the CGAS-STING1 signaling. | 0 |
| 38228616 | 2024 | Selective CK1α degraders exert antiproliferative activity against a broad range of human cancer cell lines. | 2 |
| 38254663 | 2024 | Casein Kinase 1 Phosphomimetic Mutations Negatively Impact Connexin-43 Gap Junctions in Human Pluripotent Stem Cell-Derived Cardiomyocytes. | 0 |
| 38728403 | 2024 | Substrate displacement of CK1 C-termini regulates kinase specificity. | 0 |
| 37723657 | 2024 | CSNK1A1/CK1α suppresses autoimmunity by restraining the CGAS-STING1 signaling. | 0 |
| 38228616 | 2024 | Selective CK1α degraders exert antiproliferative activity against a broad range of human cancer cell lines. | 2 |
| 38254663 | 2024 | Casein Kinase 1 Phosphomimetic Mutations Negatively Impact Connexin-43 Gap Junctions in Human Pluripotent Stem Cell-Derived Cardiomyocytes. | 0 |
| 38728403 | 2024 | Substrate displacement of CK1 C-termini regulates kinase specificity. | 0 |
| 36308624 | 2023 | Casein kinase 1α mediates eryptosis: a review. | 4 |
| 36720664 | 2023 | Overexpressions of RHOA, CSNK1A1, DVL2, FZD8, and LRP5 genes enhance gastric cancer development in the presence of Helicobacter pylori. | 0 |
| 36308624 | 2023 | Casein kinase 1α mediates eryptosis: a review. | 4 |
| 36720664 | 2023 | Overexpressions of RHOA, CSNK1A1, DVL2, FZD8, and LRP5 genes enhance gastric cancer development in the presence of Helicobacter pylori. | 0 |
| 35588152 | 2022 | Decreased Expressions of CK1α and PTEN in Sinonasal Inverted Papilloma. | 0 |
| 35985475 | 2022 | CK1α upregulates the IFNAR1 expression to prompt the anti-HBV effect of type I IFN in hepatoma carcinoma cells. | 1 |
| 35588152 | 2022 | Decreased Expressions of CK1α and PTEN in Sinonasal Inverted Papilloma. | 0 |
Citation
Max Doerfel ; Otmar Huber
CSNK1A1 (casein kinase 1, alpha 1)
Atlas Genet Cytogenet Oncol Haematol. 2008-05-01
Online version: http://atlasgeneticsoncology.org/gene/40168/csnk1a1
