FSTL3 (follistatin-like 3 (secreted glycoprotein))
2012-09-01 Michael Grusch AffiliationMedical University of Vienna, Department of Medicine I, Institute of Cancer Research, Borschkegasse 8a, A-1090 Vienna, Austria
DNA/RNA
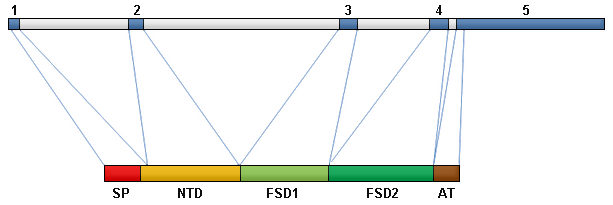
Intron/exon structure of the FSTL3 gene and domain architecture of FSTL3 protein. 1,2,3,4,5: exon number; SP: signal peptide; NTD: N-terminal domain; FSD 1: follistatin domain 1; FSD 2: follistatin domain 2; AT: acidic tail.
Description
The human FSTL3 gene is comprised of five exons spanning 7004 bp on chromosome 19p13 and gives rise to a main transcript of 2525 bp. The first exon encodes the signal peptide plus nine residues of the N-terminal domain, the second exon encodes the remainder of the N-terminal domain, the third and the fourth exon each code for a follistatin module and the fifth exon encodes the C-terminus rich in acidic amino acids (Schneyer et al., 2001; Sidis et al., 2002).
Transcription
Transcription of FSTL3 mRNA was shown to be stimulated by TGF beta and activin A via Smad proteins (Bartholin et al., 2001; Bartholin et al., 2002), which seems to be part of a negative feedback loop as FSTL3 can antagonize activin A (see below). In a different study it was found that GDF9 - another TGF beta superfamily cytokine - can suppress both basal and activin A-induced FSTL3 (and follistatin) mRNA and protein levels in cultured human granulosa-lutein (hGL) cells from women undergoing in vitro fertilization (IVF) treatment (Shi et al., 2011). In addition NF-kappaB responsive elements have been identified in the FSTL3 promoter and are involved in the cooperative stimulation of FSTL3 transcription by TNF alpha and TGF beta (Bartholin et al., 2007). Upregulation of FSTL3 mRNA was observed as a response to hypoxia in human trophoblasts and a role for hypoxia inducible factor (HIF1 alpha) was suggested. In another study a combination of 17 beta estradiol and progesterone, but not either factor alone, stimulated FSTL3 expression in cultured human endometrial stromal cells (Wang et al., 2003). An analysis of the murine FSTL3 promoter identified phorbol 12-myristate 13-acetate (PMA) but not cAMP as transcriptional activators.
Proteins
Description
The FSTL3 protein precursor consists of 263 amino acids. Amino acids 1-26 form the signal peptide, which is reponsible for directing the protein to the secretory pathway but not present in the mature secreted protein. FSTL3 has been described to contain 2 follistatin (FS) domains (aa 97-168 and 169-244) (Schneyer et al., 2001) characterized by a conserved arrangement of 10 cysteine residues. The FS domains also have similarity with the Kazal domains found in multiple serine protease inhibitors. The FSTL3 protein contains two potential N-glycosilation sites on asparagines 73 and 215. Peptide N-glycosidase F digestion reduced the size of ectopically expressed FSTL3 from 33 to 27 kDa in Western analysis (Hayette et al., 1998).
Expression
FSTL3 is expressed in a wide variety of tissues and organs with the highest expression in placenta and testis and low to absent expression in the hematopoietic system (Hayette et al., 1998; Tortoriello et al., 2001).
Localisation
The signal peptide directs the nascent protein to the secretory pathway and FSTL3 has been detected in cell culture supernatants and in human serum (Hayette et al., 1998; Hill et al., 2002; Pryor-Koishi et al., 2007). In addition to its role as a secreted protein FSTL3 has also been found in the cell nucleus in a number of different cell lines, primary cells (Tortoriello et al., 2001) and tissue sections (Torres et al., 2007). Nuclear FSTL3 was glycosylated, albeit to a lower degree than secreted FSTL3, suggesting that it enters the nucleus after partial processing in the ER (Saito et al., 2005).
Function
Similar to follistatin, secreted FSTL3 was shown to bind activin A and with lower affinity several other members of the TGF beta family including activin B, myostatin and BMP2, BMP6, and BMP7 (but excluding for instance TGF beta and BMP4) (Tsuchida et al., 2000; Tortoriello et al., 2001; Sidis et al., 2002; Hill et al., 2002; Schneyer et al., 2003). Affinity for BMPs, however, seems to be quite low and has been questioned by other studies (Sidis et al., 2006). FSTL3 bound ligands are unable to activate their cognate receptors and thus FSTL3 blocks signal transduction. Structural and biochemical data have implicated the N-terminal domain and the two FS domains in ligand binding and antagonism (Sidis et al., 2005; Stamler et al., 2008; Cash et al., 2012).
In contrast to follistatin, FSTL3 does not contain a heparin binding motif and does not bind cell surface heparan-sulfate proteoglycans (Sidis et al., 2002).
In addition to binding TGF beta family ligands, FSTL3 was demonstrated to interact with ADAM12 (a disintegrin and metalloprotease 12), an extracellular protein that has been implicated in insulin-like growth factor (IGF) and epidermal growth factor (EGF) receptor signaling (Bartholin et al., 2005), as well as with fibronectin type 1, which could indicate a role of FSTL3 in cell adhesion (Maguer-Satta et al., 2006).
Nuclear FSTL3 was shown to interact with AF10 (ALL-1 fused gene from chromosome 10) enhancing its homo-oligomerization and transcriptional activity (Forissier et al., 2007).
FSTL3 knock-out mice had increased pancreatic island number and size, enhanced insulin sensitivity and hepatic steatosis suggesting a role of FSTL3 in glucose and fat homeostasis (Mukherjee et al., 2007). Combined knockout of FSTL3 and follistatin, however, led to increased fat mass and insulin resistance despite elevated insulin production (Brown et al., 2011).
In contrast to follistatin, FSTL3 does not contain a heparin binding motif and does not bind cell surface heparan-sulfate proteoglycans (Sidis et al., 2002).
In addition to binding TGF beta family ligands, FSTL3 was demonstrated to interact with ADAM12 (a disintegrin and metalloprotease 12), an extracellular protein that has been implicated in insulin-like growth factor (IGF) and epidermal growth factor (EGF) receptor signaling (Bartholin et al., 2005), as well as with fibronectin type 1, which could indicate a role of FSTL3 in cell adhesion (Maguer-Satta et al., 2006).
Nuclear FSTL3 was shown to interact with AF10 (ALL-1 fused gene from chromosome 10) enhancing its homo-oligomerization and transcriptional activity (Forissier et al., 2007).
FSTL3 knock-out mice had increased pancreatic island number and size, enhanced insulin sensitivity and hepatic steatosis suggesting a role of FSTL3 in glucose and fat homeostasis (Mukherjee et al., 2007). Combined knockout of FSTL3 and follistatin, however, led to increased fat mass and insulin resistance despite elevated insulin production (Brown et al., 2011).
Homology
FSTL3 has a similar domain architecture as follistatin, but harbours only two instead of three follistatin modules (Tortoriello et al., 2001). Follistatin modules are found in varying numbers in a wider set of secreted proteins including FSTL1, SPARC/osteonectin, or agrin (Ullman and Perkins, 1997). With respect to activin binding ability, functional homology among follistatin domain-containing proteins has only been demonstrated for FSTL3 and follistatin (Tsuchida et al., 2000). Between different species FSTL3 is clearly conserved (96% amino acid identity between mouse and rat and about 80% for either of these species compared to human), albeit to a lesser degree than follistatin, which has around 97% amino acid identity between human, mouse and rat.
Mutations
Somatic
A t(11;19)(q13;p13) translocation has been described in a case of B-cell chronic lymphocytic leukemia. The chromosomal breakpoint occurred about 7 kbp upstream of FSTL3 leading to ectopic expression of the complete FSTL3 protein, likely as a consequence of altered upstream cis-regulatory sequences (Hayette et al., 1998). In the same study structural rearrangement of the FSTL3 locus has also been described in a case of non-Hodgkin lymphoma.
A deletion of about 1.2 Mb on chromosome 19p13.3 was identified in a patient with cleft palate, hearing impairment, congenital heart malformation, keloid scarring, immune dysregulation, and mild learning difficulties. The deleted area encompasses about 60 genes including FSTL3 (Archer et al., 2005).
A deletion of about 1.2 Mb on chromosome 19p13.3 was identified in a patient with cleft palate, hearing impairment, congenital heart malformation, keloid scarring, immune dysregulation, and mild learning difficulties. The deleted area encompasses about 60 genes including FSTL3 (Archer et al., 2005).
Implicated in
Entity name
Malignant blood disorders
Disease
Structural rearrangement of the FSTL3 locus has been described in malignant blood disorders (see above) and it has been suggested that deregulated expression of FSTL3 could contribute to leukemogenesis (Hayette et al., 1998).
Entity name
Breast carcinoma
Disease
High expression of FSTL3 was observed in invasive breast carcinoma compared to normal luminal epithelial cells. SiRNA-mediated knock-down of FSTL3 expression in breast cancer cell lines lead to growth inhibition, smad2 phosphorylation and increased transcription of activin target genes (Razanajaona et al., 2007). These observations suggest that FSTL3 may contribute to tumorigenesis by antagonizing growth limiting activin effects. In agreement with this notion another study reported increased FSTL3 mRNA and protein levels in the epithelial compartment in infiltrating ductal carcinoma of the breast when compared to normal breast tissue (Bloise et al., 2009).
Entity name
Hepatocellular carcinoma
Disease
FSTL3 expression levels were increased in hepatocytes in chemical hepatocarcinogenesis models in the rat but decreased in hepatocellular carcinoma (HCC) in humans (Grusch et al., 2006).
Entity name
Endometrial adenocarcinoma
Disease
Downregulation of FSTL3 expression was found in human endometrial adenocarcinoma (Ciarmela et al., 2004).
Entity name
Endometriosis
Disease
Ovarian endometriotic lesions showed a deranged expression of FSTL3 as well as follistatin. While follistatin expression was increased, FSTL3 mRNA and protein expression were lower in ovarian endometriosis than in healthy eutopic endometrium (Torres et al., 2007).
Entity name
Pre-eclampsia
Disease
The mRNA and protein levels of FSTL3 were compared in placentas and maternal sera of women with uncomplicated pregnancy and those with pre-eclampsia. FSTL3 was upregulated in syncytiotrophoblast cells of pre-eclamptic placental tissue and in maternal serum. Its further evaluation as potential addition to existing diagnostic markers of pre-eclampsia was suggested (Pryor-Koishi et al., 2007).
Entity name
Heart disease
Disease
FSTL3 (and FSTL1) were elevated in heart failure. FSTL3 expression correlated with markers of disease severity and returned to normal after recovery. The protein was localized to myocytes and endothelium and the expression profile of FSTL3 on microarrays revealed an association with the nuclear compartment and with genes involved in signal transduction and transcription (Lara-Pezzi et al., 2008). In a mouse model activin A and FSTL3 were upregulated in the herat by ischemic injury. While activin A protected myocytes from cell death, this effect was antagonized by FSTL3 (Oshima et al., 2009).
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|
Other Information
Locus ID:
NCBI: 10272
MIM: 605343
HGNC: 3973
Ensembl: ENSG00000070404
Variants:
dbSNP: 10272
ClinVar: 10272
TCGA: ENSG00000070404
COSMIC: FSTL3
RNA/Proteins
Expression (GTEx)
Pathways
| Pathway | Source | External ID |
|---|---|---|
| Signal Transduction | REACTOME | R-HSA-162582 |
| Signaling by Activin | REACTOME | R-HSA-1502540 |
| Antagonism of Activin by Follistatin | REACTOME | R-HSA-2473224 |
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 36502285 | 2023 | FSTL3 is highly expressed in adipose tissue of individuals with overweight or obesity and is associated with inflammation. | 3 |
| 36807490 | 2023 | The cytoplasmic expression of FSTL3 correlates with colorectal cancer progression, metastasis status and prognosis. | 1 |
| 37739334 | 2023 | Follistatin and follistatin-like 3 in metabolic disorders. | 0 |
| 37751246 | 2023 | Prognostic Impact of FSTL3, ADAM12, and FAT4 in Patients of Colon Cancer: Clinicopathologic Study. | 0 |
| 37904173 | 2023 | FSTL3 partially mediates the association of increased nonalcoholic fatty liver disease fibrosis risk with acute myocardial infarction in patients with type 2 diabetes mellitus. | 1 |
| 38044354 | 2023 | Stroma-associated FSTL3 is a factor of calcium channel-derived tumor fibrosis. | 0 |
| 36502285 | 2023 | FSTL3 is highly expressed in adipose tissue of individuals with overweight or obesity and is associated with inflammation. | 3 |
| 36807490 | 2023 | The cytoplasmic expression of FSTL3 correlates with colorectal cancer progression, metastasis status and prognosis. | 1 |
| 37739334 | 2023 | Follistatin and follistatin-like 3 in metabolic disorders. | 0 |
| 37751246 | 2023 | Prognostic Impact of FSTL3, ADAM12, and FAT4 in Patients of Colon Cancer: Clinicopathologic Study. | 0 |
| 37904173 | 2023 | FSTL3 partially mediates the association of increased nonalcoholic fatty liver disease fibrosis risk with acute myocardial infarction in patients with type 2 diabetes mellitus. | 1 |
| 38044354 | 2023 | Stroma-associated FSTL3 is a factor of calcium channel-derived tumor fibrosis. | 0 |
| 34335633 | 2021 | Follistatin-Like 3 Correlates With Lymph Node Metastasis and Serves as a Biomarker of Extracellular Matrix Remodeling in Colorectal Cancer. | 15 |
| 34425560 | 2021 | Regulation of follistatin-like 3 expression by miR-486-5p modulates gastric cancer cell proliferation, migration and tumor progression. | 6 |
| 34335633 | 2021 | Follistatin-Like 3 Correlates With Lymph Node Metastasis and Serves as a Biomarker of Extracellular Matrix Remodeling in Colorectal Cancer. | 15 |
Citation
Michael Grusch
FSTL3 (follistatin-like 3 (secreted glycoprotein))
Atlas Genet Cytogenet Oncol Haematol. 2012-09-01
Online version: http://atlasgeneticsoncology.org/gene/111/fstl3-(follistatin-like-3-(secreted-glycoprotein))
Historical Card
2009-01-01 FSTL3 (follistatin-like 3 (secreted glycoprotein)) by Michael Grusch Affiliation
Medical University of Vienna, Department of Medicine I, Institute of Cancer Research, Borschkegasse 8a, A-1090 Vienna, Austria
