KLK4 (kallikrein-related peptidase 4)
2009-01-01 John Lai , Ying Dong , Judith A Clements AffiliationHormone Dependent Cancer Program, Institute of Health, Biomedical Innovation (IHBI), Queensland University of Technology (QUT), Brisbane, Australia
Identity
HGNC
LOCATION
19q13.41
LOCUSID
ALIAS
AI2A1,ARM1,EMSP,EMSP1,KLK-L1,PRSS17,PSTS,kallikrein
FUSION GENES
DNA/RNA
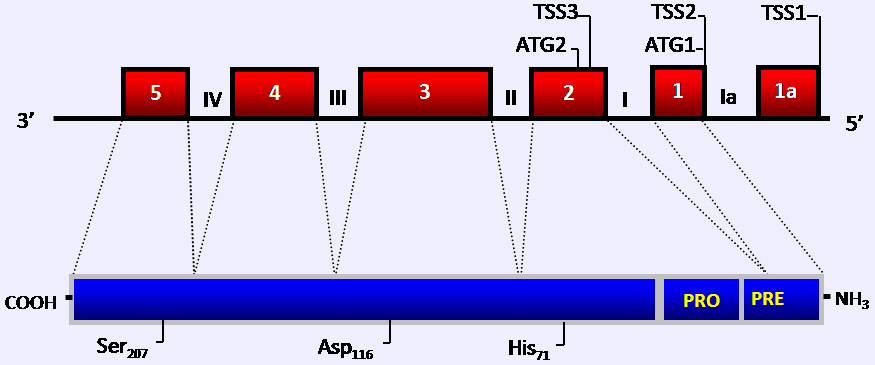
Genomic and protein structure of the KLK4 gene. The KLK4 gene is classically comprised of 5 exons (grey boxes, classic numerals) and 4 introns (roman numerals), although extra 5 UTR sequences in exon 1a have also been described. Shown here are the three putative alternative transcription start sites (TSS1, TSS2 and TSS3). Also shown are the postions of ATG1 and ATG2 that would be utilised from these variant transcripts. The amino acid numbering for the residues of the catalytic triad (His71, Asp116, Ser207) are relative to the full-length protein starting from ATG1.
Description
The gene encompasses 4.38 kb of gDNA.
Transcription
Three alternative transcription start sites (TSSs) have been predicted or identified experimentally for the KLK4 gene. TSS1 was identified using 5 RACE and is located 639 bp upstream of TSS2, which was predicted using in silico analysis. TSS3 was also identified using 5 RACE and is located 1396 bp downstream of TSS1.
Nine KLK4 variant transcripts have been identified. These variants include the TSS variants, variants with 5 UTR deletion for TSS1 and TSS2 transcripts, exon 4 deletion and partial intron II (12bp) insertion variants, intron 3 insertion variants, and variants with combinations thereof.
Nine KLK4 variant transcripts have been identified. These variants include the TSS variants, variants with 5 UTR deletion for TSS1 and TSS2 transcripts, exon 4 deletion and partial intron II (12bp) insertion variants, intron 3 insertion variants, and variants with combinations thereof.
Pseudogene
Not identified.
Proteins
Description
Two KLK4 protein isoforms, full-length KLK4 (254 amino acids) and an N-terminal truncated KLK4 (205 amino acids) have been described. Full-length KLK4 has a secretion signal (pre-) peptide (26 amino acids), followed by an activation (pro-) peptide (4 amino acids) and the mature chain (224 amino acids) with 1 potential N-linked glycosylation site. The catalytic triad of His71, Asp116, Ser207 (relative to Met = 1 and encoded by ATG1 in exon 1) is conserved and is essential for proteolytic activity. After synthesis as a KLK4 full-length protein, the signal peptide is then cleaved and pro-KLK4 (zymogen) is subsequently secreted from the cell. Upon activation, the propeptide is removed to generate the mature active enzyme. KLK4 complexes with alpha(1)-antitrypsin and alpha(2)-antiplasmin and alpha(2)-macroglobulin.
The N-terminal truncated KLK4 isoform initiated from the ATG2 in exon 2, has the pre-pro-region and 9 amino acids from the mature KLK4 omitted, although the catalytic triad remains. Further, the truncated KLK4 isoform is not glycosylated despite retaining the potential N-linked glycosylation site.
The N-terminal truncated KLK4 isoform initiated from the ATG2 in exon 2, has the pre-pro-region and 9 amino acids from the mature KLK4 omitted, although the catalytic triad remains. Further, the truncated KLK4 isoform is not glycosylated despite retaining the potential N-linked glycosylation site.
Expression
The KLK4 gene was originally designated the PRSS17 gene and was cloned from the cells of the enamel organ epithelia of developing teeth in pig using degenerate primers encoding the EMPS1 protein. Subsequent studies in human tissues using Northern blot analysis have shown that KLK4 mRNA expression is predominantly localised to the prostate, although more sensitive RT-PCR experiments have shown that the breast, ovary, endometrium, salivary gland, lung, adrenal gland, colon, trachea, brain, testis, spinal cord, thyroid, skin and kidney also express KLK4 mRNA at low to modest levels. KLK4 mRNA has also been detected in sebaceous glands, sweat glands, hair follicles, stratum basale, stratum spinosum and stratum granulosum by in situ hybridisation experiments. In pig, KLK4 mRNA is expressed in the endometrium in the early stages of the oestrous cycle.
KLK4 protein has been detected in a wide range of tissues at low (adrenal, aorta, brain, breast, cervix, heart, kidney, liver, muscle, pancreas, pituitary, salivary gland, small intestine, spinal cord, spleen, testis, skin, thyroid, and uterus) to high (prostate) levels. Modest levels of KLK4 protein have also been detected in body fluid, such as seminal plasma and urine. High KLK4 levels have also been detected in the prostate, breast and ovarian cancer tissues from patients.
KLK4 protein has been detected in a wide range of tissues at low (adrenal, aorta, brain, breast, cervix, heart, kidney, liver, muscle, pancreas, pituitary, salivary gland, small intestine, spinal cord, spleen, testis, skin, thyroid, and uterus) to high (prostate) levels. Modest levels of KLK4 protein have also been detected in body fluid, such as seminal plasma and urine. High KLK4 levels have also been detected in the prostate, breast and ovarian cancer tissues from patients.
Localisation
Full-length KLK4 encodes a secreted protein and is localised intracellularly to the cytoplasm, although GFP labelled N-terminal truncated isoforms have been found to be predominantly localised to the nucleus.
Function
To date, the major biological function of KLK4 has been derived from porcine studies which have shown that KLK4 is important in dental enamel mineralisation by degrading amelogenin, the major protein in the enamel matrix of developing teeth. More recently, KLK4 was also reported to degrade porcine enamelin which is another protein found in developing teeth. Further, in mice, KLK4 was only expressed by transition and maturation stage ameloblasts, which is consistent with KLK4 functioning primarily to degrade the enamel matrix in developing teeth.
The role of KLK4 in the prostate, a tissue that highly expresses KLK4, is less clearly defined although it is thought to be important in prostate cancer progression given its role in degrading extracellular matrix (ECM) proteins in teeth, and potentially increasing IGF levels by degrading IGFBP3, IGFBP4, IGFBP5, IGFBP6. KLK4 is also reported to activate pro- HGFA and thereby potentially leading to tumour progression through activation of the MET receptor. A substrate specificity screening study has shown that full-length KLK4 has trypsin-like specificity and potentially activates proteins, such as, prostate specific antigen (PSA) / KLK3, bone morphogenetic protein (BMP) family and parathyroid hormone-related protein ( PTHrP ) that are involved in normal and neoplastic prostate (patho)-physiology. Recombinant KLK4 was also reported to be a better activator of PSA and single chain urokinase-type plasminogen activator than KLK2. More recently, KLK4 was shown to activate other members of the kallikrein-related peptidase family including KLK1, KLK2, KLK3, KLK5, KLK6, KLK9, KLK11, KLK12, KLK13, KLK14, KLK15. However, the precise physiological role for KLK4 in the prostate and other tissues remains to be identified. The function of the N-terminal truncated KLK4 remains to be established.
The role of KLK4 in the prostate, a tissue that highly expresses KLK4, is less clearly defined although it is thought to be important in prostate cancer progression given its role in degrading extracellular matrix (ECM) proteins in teeth, and potentially increasing IGF levels by degrading IGFBP3, IGFBP4, IGFBP5, IGFBP6. KLK4 is also reported to activate pro- HGFA and thereby potentially leading to tumour progression through activation of the MET receptor. A substrate specificity screening study has shown that full-length KLK4 has trypsin-like specificity and potentially activates proteins, such as, prostate specific antigen (PSA) / KLK3, bone morphogenetic protein (BMP) family and parathyroid hormone-related protein ( PTHrP ) that are involved in normal and neoplastic prostate (patho)-physiology. Recombinant KLK4 was also reported to be a better activator of PSA and single chain urokinase-type plasminogen activator than KLK2. More recently, KLK4 was shown to activate other members of the kallikrein-related peptidase family including KLK1, KLK2, KLK3, KLK5, KLK6, KLK9, KLK11, KLK12, KLK13, KLK14, KLK15. However, the precise physiological role for KLK4 in the prostate and other tissues remains to be identified. The function of the N-terminal truncated KLK4 remains to be established.
Homology
At the protein level, KLK4 shares 25%, (KLK12), 29% (KLK10), 31% (KLK9), 34% (KLK1, 2, 3), 35% (KLK8, 13), 36% (KLK6), 37% (KLK15), 40% (KLK14), 43% (KLK7), 45% (KLK5) sequence homology with other members of the kallikrein-related peptidase family. Unlike KLK1, KLK2 and KLK3, KLK4 lacks the "kallikrein loop", a region of 11 amino acids encoded in exon 3 and thought to be important in the substrate specificity of these enzymes. KLK4 shares 72% sequence homology to the porcine EMPS1 protein. Bayesian phylogenetic analyses suggests that KLK4, KLK5 and KLK14 may form a sub-family within the kallikrein-related peptidase gene family.
Mutations
Germinal
A mutation in the KLK4 gene (G.2142>A) that encodes for a truncated KLK4 protein that lacks S207, which is necessary for catalytic activity, has been shown to associate with autosomal recessive hypomaturation amelogenesis imperfecta, which is a disorder affecting tooth enamel formation. Comparison of tooth enamel in patients with the G- and A-alleles suggest that wild-type KLK4 (G allele) is important for the final removal of the extracellar enamel matrix proteins for normal enamel maturation.
Implicated in
Entity name
Hormone dependent cancers
Note
It has been proposed that like PSA, encoded by the KLK3 gene, the KLK4 gene may play a role in hormone dependent cancers given its (a) higher expression in endocrine cells (b) regulation by hormones such as androgens, oestradiol and progestins (c) dysregulated expression in cancer cells and (d) potential role in extra-cellular matrix degradation and growth factor activation.
Entity name
Prostate cancer
Note
Hormone dependent cancer
Disease
Kallikrein 4 has been reported to be more highly expressed in cancerous than benign prostate tissues at both the mRNA and protein levels. For example, a tissue microarray study carried out on 42 benign and 207 malignant prostate tissues found that KLK4 was more highly expressed in prostate cancer cells when compared to benign cells, although one study using sandwich-type immunoassay on 16 malignant and 18 benign prostate tissues found no difference in KLK4 expression. It has also been shown that KLK4 specific antibodies can be detected in sera from prostate cancer patients. In prostate cancer cells, KLK4 has been shown to be up-regulated by androgens at both the mRNA and protein level. In two expression studies using Northern blot analysis, KLK4 mRNA expression was found to be up-regulated by 18- and 1.7 -fold after treatment of LNCaP cells with R1881 for 24 and 48 hr, respectively. Further, prostate cancer cells (PC-3 and DU145) transfected with KLK4 have also been shown to have increased cellular migration, proliferation and colony formation. Over-expression of KLK4 in prostate cancer cells has also been associated with increased migration of these cells to SaOs2 conditioned media and greater attachment to bone matrix proteins collagens I and IV. The role of KLK4 in prostate biology is thought to be mediated in part through activation of the PAR-1/F2R and PAR-2/F2RL1 signalling pathways, although the precise mechanisms and importance in prostate cancer remain to be identified.
Cytogenetics
Comparison of two human prostate cell lines, P69SV40Tag (P69) and its tumorigenic subline, M12, and 11 prostate cancer cases showed LOH in M12 at 19q13.42, which is proximal to the KLK4 locus.
Hybrid gene
No KLK4 fusion transcript or protein has been reported thus far.
Entity name
Epithelial ovarian carcinoma
Note
Hormone dependent cancer
Disease
Kallikrein 4 is more highly expressed in serous ovarian carcinomas at both the mRNA and protein levels, and high KLK4 mRNA expression is associated with poorly differentiated and late clinical stage ovarian carcinomas. KLK4 mRNA was also found to be expressed in tumour cells isolated from ascites fluid in 60% (6/10) of ovarian cancer patients. KLK4 protein expression was also found to be more highly expressed in paclitaxel-resistant (79/93; 85%) than paclitaxel-sensitive tumours (20/33; 61%), suggesting that KLK4 may have utility as a predictive marker for chemoresistant ovarian cancers. Co-overexpression of KLK4, 5, 6 and 7 in ovarian cancer cells (OV-MZ-6) led to increased invasion in vitro and simultaneous expression of these KLKs in nude mice resulted in increased tumour burden.
Prognosis
KLK4 mRNA expression in tumour tissues indicates shorter overall and progression free survival time for epithelial ovarian carcinoma patients. It is also an independent indicator of poor prognosis in patients with grade 1 and 2 tumours.
Entity name
Breast cancer
Note
Hormone dependent cancer
Disease
Comparison of RNA levels from normal and malignant breast tissue using real time RT-PCR showed that KLK4 expression was up-regulated in cancer cells. Further analysis using laser dissection of these tumours and immuno-histochemistry showed that the observed increase of KLK4 expression was due to increased expression in the surrounding stromal cells.
Entity name
Endometrial cancer
Note
Hormone dependent cancer
Disease
KLK4 protein levels were shown by Western blot analysis to be up-regulated in endometrial KLE cells in response to both oestradiol and progestin. This response was increased when cells were simultaneously treated with oestrogen and progesterone.
Entity name
Non-small cell lung cancer (NSCLC)
Disease
Using a KLK4 ELISA on 51 patients with NSCLC and 50 normal controls, it was shown that KLK4 may have utility as a lung cancer biomarker when used in conjunction with KLK8, KLK10, KLK11, KLK12, KLK13, and KLK14.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 18499550 | 2008 | The genetic basis of inherited anomalies of the teeth. Part 1: clinical and molecular aspects of non-syndromic dental disorders. | Bailleul-Forestier I et al |
| 17976015 | 2007 | Defining the extended substrate specificity of kallikrein 1-related peptidases. | Borgoño CA et al |
| 18627289 | 2008 | Kallikreins as microRNA targets: an in silico and experimental-based analysis. | Chow TF et al |
| 11258672 | 2001 | The expanded human kallikrein (KLK) gene family: genomic organisation, tissue-specific expression and potential functions. | Clements J et al |
| 12370833 | 2002 | Characterization of KLK4 expression and detection of KLK4-specific antibody in prostate cancer patient sera. | Day CH et al |
| 18627343 | 2008 | Structures and specificity of the human kallikrein-related peptidases KLK 4, 5, 6, and 7. | Debela M et al |
| 16322328 | 2005 | Compartmentalized expression of kallikrein 4 (KLK4/hK4) isoforms in prostate cancer: nuclear, cytoplasmic and secreted forms. | Dong Y et al |
| 11489814 | 2001 | Human kallikrein 4 (KLK4) is highly expressed in serous ovarian carcinomas. | Dong Y et al |
| 16829021 | 2006 | In silico identification and Bayesian phylogenetic analysis of multiple new mammalian kallikrein gene families. | Elliott MB et al |
| 17127754 | 2006 | Porcine endometrial and conceptus tissue kallikrein 1, 4, 11, and 14 gene expression. | Fernando SC et al |
| 11054574 | 2000 | Sequencing and expression analysis of the serine protease gene cluster located in chromosome 19q13 region. | Gan L et al |
| 17221837 | 2007 | Kallikrein 4 is a potential mediator of cellular interactions between cancer cells and osteoblasts in metastatic prostate cancer. | Gao J et al |
| 14684688 | 2003 | Novel ENAM mutation responsible for autosomal recessive amelogenesis imperfecta and localised enamel defects. | Hart TC et al |
| 10690663 | 2000 | Localization of EMSP1 expression during tooth formation and cloning of mouse cDNA. | Hu JC et al |
| 17552940 | 2007 | Developmental biology and genetics of dental malformations. | Hu JC et al |
| 12206593 | 2002 | Enamelysin and kallikrein-4 mRNA expression in developing mouse molars. | Hu JC et al |
| 16612087 | 2005 | Proteomics and genetics of dental enamel. | Hu JC et al |
| 10863090 | 2000 | Characterization of the mouse and human PRSS17 genes, their relationship to other serine proteases, and the expression of PRSS17 in developing mouse incisors. | Hu JC et al |
| 12077288 | 2002 | Identification of naturally processed CD4 T cell epitopes from the prostate-specific antigen kallikrein 4 using peptide-based in vitro stimulation. | Hural JA et al |
| 17545602 | 2007 | Kallikrein 4 is a proliferative factor that is overexpressed in prostate cancer. | Klokk TI et al |
| 12925213 | 2003 | Expression and localization of tissue kallikrein mRNAs in human epidermis and appendages. | Komatsu N et al |
| 11506707 | 2001 | Distinctly different gene structure of KLK4/KLK-L1/prostase/ARM1 compared with other members of the kallikrein family: intracellular localization, alternative cDNA forms, and Regulation by multiple hormones. | Korkmaz KS et al |
| 17587816 | 2007 | Epithelial-mesenchymal transition in prostate cancer and the potential role of kallikrein serine proteases. | Lawrence MG et al |
| 18484629 | 2008 | Mutational spectrum of FAM83H: the C-terminal portion is required for tooth enamel calcification. | Lee SK et al |
| 18627287 | 2008 | Functions of KLK4 and MMP-20 in dental enamel formation. | Lu Y et al |
| 18687310 | 2008 | Specific increase of human kallikrein 4 mRNA and protein levels in breast cancer stromal cells. | Mangé A et al |
| 15389820 | 2005 | Substrates of the prostate-specific serine protease prostase/KLK4 defined by positional-scanning peptide libraries. | Matsumura M et al |
| 15650036 | 2005 | Intron retention: a common splicing event within the human kallikrein gene family. | Michael IP et al |
| 18221492 | 2008 | Activation of hepatocyte growth factor activator zymogen (pro-HGFA) by human kallikrein 1-related peptidases. | Mukai S et al |
| 11344246 | 2001 | Kallikrein 4 (KLK4), a new member of the human kallikrein gene family is up-regulated by estrogen and progesterone in the human endometrial cancer cell line, KLE. | Myers SA et al |
| 14630899 | 2003 | Relative levels of mRNA encoding enamel proteins in enamel organ epithelia and odontoblasts. | Nagano T et al |
| 10077646 | 1999 | Molecular cloning and characterization of prostase, an androgen-regulated serine protease with prostate-restricted expression. | Nelson PS et al |
| 15911097 | 2005 | Human tissue kallikrein gene family: applications in cancer. | Obiezu CV et al |
| 15961548 | 2005 | Human kallikrein 4: quantitative study in tissues and evidence for its secretion into biological fluids. | Obiezu CV et al |
| 16557045 | 2006 | Kallikreins as markers of disseminated tumour cells in ovarian cancer-- a pilot study. | Oikonomopoulou K et al |
| 12437987 | 2002 | Organization and evolution of the glandular kallikrein locus in Mus musculus. | Olsson AY et al |
| 18096894 | 2008 | Premature stop codon in MMP20 causing amelogenesis imperfecta. | Papagerakis P et al |
| 17125728 | 2007 | Phenotype and enamel ultrastructure characteristics in patients with ENAM gene mutations g.13185-13186insAG and 8344delG. | Pavlic A et al |
| 18316555 | 2008 | A multiparametric serum kallikrein panel for diagnosis of non-small cell lung carcinoma. | Planque C et al |
| 16800744 | 2006 | Overexpression of the human tissue kallikrein genes KLK4, 5, 6, and 7 increases the malignant phenotype of ovarian cancer cells. | Prezas P et al |
| 18308730 | 2008 | Kallikrein-related peptidase 4 (KLK4) initiates intracellular signaling via protease-activated receptors (PARs). KLK4 and PAR-2 are co-expressed during prostate cancer progression. | Ramsay AJ et al |
| 12664466 | 2002 | Porcine kallikrein-4 activation, glycosylation, activity, and expression in prokaryotic and eukaryotic hosts. | Ryu O et al |
| 17266769 | 2007 | Exclusion of known gene for enamel development in two Brazilian families with amelogenesis imperfecta. | Santos MC et al |
| 11062993 | 1998 | Enamel matrix serine proteinase 1: stage-specific expression and molecular modeling. | Scully JL et al |
| 9465170 | 1998 | Purification, characterization, and cloning of enamel matrix serine proteinase 1. | Simmer JP et al |
| 12489196 | 2002 | Expression, structure, and function of enamel proteinases. | Simmer JP et al |
| 16304440 | 2005 | Genes and related proteins involved in amelogenesis imperfecta. | Stephanopoulos G et al |
| 10438493 | 1999 | Localization of a new prostate-specific antigen-related serine protease gene, KLK4, is evidence for an expanded human kallikrein gene family cluster on chromosome 19q13.3-13.4. | Stephenson SA et al |
| 19029081 | 2008 | Enamel proteases reduce amelogenin-apatite binding. | Sun Z et al |
| 11735417 | 2001 | Characterization of hK4 (prostase), a prostate-specific serine protease: activation of the precursor of prostate specific antigen (pro-PSA) and single-chain urokinase-type plasminogen activator and degradation of prostatic acid phosphatase. | Takayama TK et al |
| 16172196 | 2005 | Kallikrein 4 (hK4) and prostate-specific antigen (PSA) are associated with the loss of E-cadherin and an epithelial-mesenchymal transition (EMT)-like effect in prostate cancer cells. | Veveris-Lowe TL et al |
| 17629198 | 2007 | [Exclusion of candidate genes in a family with amelogenesis imperfecta]. | Wang XJ et al |
| 16800731 | 2006 | The role of kallikrein-related peptidases in prostate cancer: potential involvement in an epithelial to mesenchymal transition. | Whitbread AK et al |
| 18714142 | 2009 | Human and mouse enamel phenotypes resulting from mutation or altered expression of AMEL, ENAM, MMP20 and KLK4. | Wright JT et al |
| 16838342 | 2006 | The molecular etiologies and associated phenotypes of amelogenesis imperfecta. | Wright JT et al |
| 15262123 | 2004 | Kallikrein 4 is associated with paclitaxel resistance in ovarian cancer. | Xi Z et al |
| 15059887 | 2004 | Kallikrein 4 is a predominantly nuclear protein and is overexpressed in prostate cancer. | Xi Z et al |
| 16674662 | 2006 | How do enamelysin and kallikrein 4 process the 32-kDa enamelin? | Yamakoshi Y et al |
| 17823117 | 2007 | Activation profiles and regulatory cascades of the human kallikrein-related peptidases. | Yoon H et al |
| 10485467 | 1999 | Prostase/KLK-L1 is a new member of the human kallikrein gene family, is expressed in prostate and breast tissues, and is hormonally regulated. | Yousef GM et al |
Other Information
Locus ID:
NCBI: 9622
MIM: 603767
HGNC: 6365
Ensembl: ENSG00000167749
Variants:
dbSNP: 9622
ClinVar: 9622
TCGA: ENSG00000167749
COSMIC: KLK4
RNA/Proteins
Expression (GTEx)
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 37918799 | 2024 | Kallikrein-Related Peptidase 4 Promotes Proliferation, Migration, Invasion, and Pro-Angiogenesis of Endometrial Stromal Cells via Regulation of Brain-Derived Neurotrophic Factor Production in Endometriosis. | 0 |
| 37918799 | 2024 | Kallikrein-Related Peptidase 4 Promotes Proliferation, Migration, Invasion, and Pro-Angiogenesis of Endometrial Stromal Cells via Regulation of Brain-Derived Neurotrophic Factor Production in Endometriosis. | 0 |
| 35982181 | 2023 | Loss of KLK4::KLKP1 pseudogene expression by RNA chromogenic in-situ hybridization is associated with PTEN loss and increased risk of biochemical recurrence in a cohort of middle eastern men with prostate cancer. | 0 |
| 35982181 | 2023 | Loss of KLK4::KLKP1 pseudogene expression by RNA chromogenic in-situ hybridization is associated with PTEN loss and increased risk of biochemical recurrence in a cohort of middle eastern men with prostate cancer. | 0 |
| 36260980 | 2022 | KLK4 Silencing Inhibits the Growth of Chromophobe Renal Cell Carcinoma through ERK/AKT Signaling Pathway. | 1 |
| 36260980 | 2022 | KLK4 Silencing Inhibits the Growth of Chromophobe Renal Cell Carcinoma through ERK/AKT Signaling Pathway. | 1 |
| 34101307 | 2021 | lncRNA IGF2-AS promotes the osteogenic differentiation of bone marrow mesenchymal stem cells by sponging miR-3,126-5p to upregulate KLK4. | 4 |
| 34101307 | 2021 | lncRNA IGF2-AS promotes the osteogenic differentiation of bone marrow mesenchymal stem cells by sponging miR-3,126-5p to upregulate KLK4. | 4 |
| 31265790 | 2020 | MicroRNA-378-3p/5p suppresses the migration and invasiveness of oral squamous carcinoma cells by inhibiting KLK4 expression. | 6 |
| 31829025 | 2020 | MiR-378a-5p inhibits angiogenesis of oral squamous cell carcinoma by targeting KLK4. | 11 |
| 31868942 | 2020 | MicroRNA-378-3p/5p represses proliferation and induces apoptosis of oral squamous carcinoma cells via targeting KLK4. | 3 |
| 31265790 | 2020 | MicroRNA-378-3p/5p suppresses the migration and invasiveness of oral squamous carcinoma cells by inhibiting KLK4 expression. | 6 |
| 31829025 | 2020 | MiR-378a-5p inhibits angiogenesis of oral squamous cell carcinoma by targeting KLK4. | 11 |
| 31868942 | 2020 | MicroRNA-378-3p/5p represses proliferation and induces apoptosis of oral squamous carcinoma cells via targeting KLK4. | 3 |
| 30644845 | 2019 | Crystal structures of the complex of a kallikrein inhibitor from Bauhinia bauhinioides with trypsin and modeling of kallikrein complexes. | 2 |
Citation
John Lai ; Ying Dong ; Judith A Clements
KLK4 (kallikrein-related peptidase 4)
Atlas Genet Cytogenet Oncol Haematol. 2009-01-01
Online version: http://atlasgeneticsoncology.org/gene/41084/klk4
