CDKN2B (cyclin-dependent kinase inhibitor 2B (p15, inhibits CDK4))
2011-01-01 Joanna Fares , Linda Wolff , Juraj Bies AffiliationLab Cell Oncology, National Cancer Institute NIH, 37 Convent Dr, Bethesda MD 20892, USA (JF, LW, JB); Biochemistry, Molecular Biology Department, Georgetown University, Washington DC 20037, USA (JF)
Identity
DNA/RNA
Description
Transcription
p15: 3.82 Kb of mRNA.
p10: 0.86 Kb of mRNA.
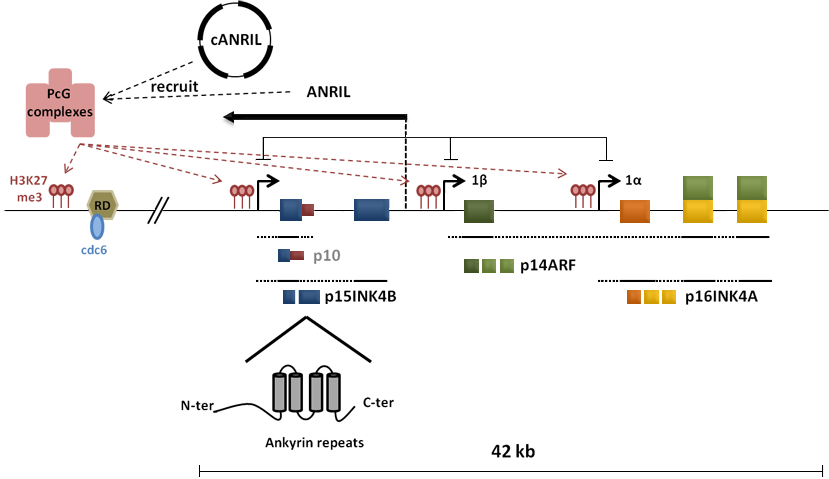
Regulation: A conserved DNA element with the ability to regulate the entire INK4/ARF locus has been identified in close proximity of the locus and named regulatory domain (RD). It appears to promote transcriptional repression of all three genes encoded by the locus, in a manner dependent on CDC6. In proliferating embryonic fibroblasts (MEFs), EZH2 a member of the polycomb repressive complex 2 (PRC2) as well as BMI1 and M33 members of the polycomb repressive complex 1 (PRC1) are strongly expressed and are found to localize to the INK4/ARF RD. BMI1 has been shown to interact specifically with CDC6. These polycomb group (PcG) complexes repress the locus activity through the establishment of repressive chromatin modifications such as H3K27 trimethylation. During senescence, binding of these complexes to RD is lost and correlates with increased expression of the INK4/ARF genes (Figure 1).
The p15INK4B gene is also silenced by a long non coding RNA, called antisense non-coding RNA in the INK4 locus (ANRIL), whose expression was found to be inversed to the expression of p15INK4B in leukemia cell lines. It was shown that ANRIL induces the silencing of p15INK4B in cis and trans by triggering heterochromatin formation in a Dicer-independent manner. PcG complexes are recruited to the INK4/ARF locus by ANRIL and modulate its repression (Figure 1). Additionally, a naturally occurring antisense circular ANRIL RNAs (cANRIL) has also been described. Different forms of cANRIL are produced in most INK4/ARF expressing cells, suggesting that alternative splicing events leading to different ANRIL structures can contribute to changes in PcG-mediated INK4/ARF repression.
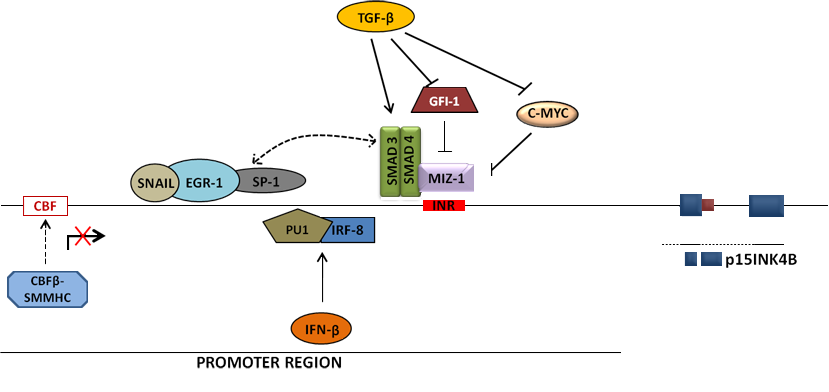
Specific transcription regulators of p15INK4B have also been reported (see Figure 2). These include TGF-b, MIZ-1, SMAD3/SMAD4 complex, SP1, c-MYC, IRF8, PU.1, SNAIL and EGR1 factors among others.
p15INK4B expression is dramatically induced by TGF-b, suggesting that it is a potent downstream effector of TGF-b mediated growth arrest.
MIZ-1 is a transcription factor that has been shown to bind the initiator element (Inr) in the promoter region and induce the transcription of p15INK4B in epithelial cells. However it can also recruit transcriptional co-repressors such as c-MYC and GFI-1 to the promoter region by binding to them and forming inhibitory complexes. TGF-b has been reported to re-activate the core promoter through downregulation of C-MYC and GFI-1, thereby releasing endogenous MIZ-1 from inhibition.
The SMAD3/4 complex readily forms following TGF-b treatment and physical interaction between this complex, MIZ-1 and promoter-bound SP1 protein has been described. These interactions have been proposed to constitute a platform for the recruitment of co-activators, and do not seem to be affected by the suppressor activity of c-MYC. The inhibitory function of c-MYC seems to be cell-type dependent, as it was confirmed in epithelial cells but not in the hematopoietic lineage. In myeloid cells, the transcription factor c-MYB was shown to prevent the transcription and the upregulation of p15INK4B which is normally associated with the differentiation process. The mechanism by which C-MYB does this is unclear but it is not through upregulation of c-MYC, a known target of c-MYB.
A tri-component transcriptional complex consisting of SNAIL, SP1 and EGR-1 was also described for its ability to trigger the p15INK4b promoter activation upon TPA treatment.
In murine myeloid cells specifically, the interferon consensus sequence-binding protein/interferon regulatory factor 8 (ICSBP/IRF-8) in combination with PU.1 were shown to bind p15Ink4b promoter and activate the transcription of the gene in response to IFN-b treatment.
In AML patients with inv(16), p15INK4B silencing was found to be caused by inv(16)-encoded core binding factor beta-smooth muscle myosin heavy chain (CBFb-SMMHC). CBFb-SMMHC was shown to displace RUNX1 from a newly determined CBF site in the promoter of p15INK4B.
Proteins
Description
p10 transcript encodes the shorter variant. The protein consists of 78 aa only and its mass is 10 KDa. It shares a similar NH2 terminus to p15 but contains a different basic COOH terminus that is translated from the p15Ink4b intronic region (Figure 2).
Expression
Localisation
Function
Three other members of the INK4 family of CDK inhibitors: p16INK4A, p18INK4C and p19INK4D are encoded by unique genes and share roughly 40% homology. They have similar protein structure characterized by the presence of four ankyrin-like motif tandem repeats that are predicted to be engaged in protein-protein interactions.
II- Function during hematopoietic cell differentiation. Another role for p15INK4B during differentiation of early hematopoietic progenitors has also been described. In knockout mice, loss of p15INK4B was shown to favor the differentiation of common myeloid progenitors (CMP) into granulocyte macrophage progenitors (GMP) resulting in an imbalance between the myeloid and the erythroid compartments.
III- Function during cellular senescence. Cellular senescence is accompanied by hallmark features that include the up-regulation of cell cycle inhibitors like p15INK4B, p16INK4A and p21CIP. When overexpressed, p15INK4B engages the RB pathway to promote a stable senescent state which has been shown to occur in part through a process that involves alterations in heterochromatin and the stable silencing of E2F target genes.
Another mechanism that has been described is the inactivation of c-MYC which results in the induction of p15INK4B expression and correlates with the global changes in heterochromatin structure known to be associated with cellular senescence.
Homology
Mutations
Note
Implicated in
This disease is also commonly referred to as B-cell chronic lymphocytic leukemia (B-CLL).
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 16343268 | 2006 | Promoter hypermethylation of p15INK4B, HIC1, CDH1, and ER is frequent in myelodysplastic syndrome and predicts poor prognosis in early-stage patients. | Aggerholm A et al |
| 11331894 | 2001 | Integrating Myc and TGF-beta signalling in cell-cycle control. | Amati B et al |
| 10764143 | 2000 | Methylation status of the p15INK4B gene in hematopoietic progenitors and peripheral blood cells in myelodysplastic syndromes. | Aoki E et al |
| 19164764 | 2009 | Gfi-1 represses CDKN2B encoding p15INK4B through interaction with Miz-1. | Basu S et al |
| 20457873 | 2010 | Myeloid-specific inactivation of p15Ink4b results in monocytosis and predisposition to myeloid leukemia. | Bies J et al |
| 21151960 | 2010 | Expression of linear and novel circular forms of an INK4/ARF-associated non-coding RNA correlates with atherosclerosis risk. | Burd CE et al |
| 9639410 | 1998 | Review of alterations of the cyclin-dependent kinase inhibitor INK4 family genes p15, p16, p18 and p19 in human leukemia-lymphoma cells. | Drexler HG et al |
| 10763830 | 2000 | Translation of p15.5INK4B, an N-terminally extended and fully active form of p15INK4B, is initiated from an upstream GUG codon. | Fuxe J et al |
| 20121949 | 2010 | Snail associates with EGR-1 and SP-1 to upregulate transcriptional activation of p15INK4b. | Hu CT et al |
| 11124804 | 2000 | Structural basis of inhibition of CDK-cyclin complexes by INK4 inhibitors. | Jeffrey PD et al |
| 7987828 | 1994 | Deletion of p16 and p15 genes in brain tumors. | Jen J et al |
| 21151178 | 2011 | Long non-coding RNA ANRIL is required for the PRC2 recruitment to and silencing of p15(INK4B) tumor suppressor gene. | Kotake Y et al |
| 10880462 | 2000 | Limited overlapping roles of P15(INK4b) and P18(INK4c) cell cycle inhibitors in proliferation and tumorigenesis. | Latres E et al |
| 11076015 | 2000 | Genetic analysis of mammalian cyclin-dependent kinases and their inhibitors. | Malumbres M et al |
| 17283131 | 2007 | Methylation-independent silencing of the tumor suppressor INK4b (p15) by CBFbeta-SMMHC in acute myelogenous leukemia with inv(16). | Markus J et al |
| 12809602 | 2003 | Rb-mediated heterochromatin formation and silencing of E2F target genes during cellular senescence. | Narita M et al |
| 17546638 | 2007 | Hypermethylation of the p15INK4B gene promoter in B-chronic lymphocytic leukemia. | Papageorgiou SG et al |
| 18985067 | 2008 | An INKlination for epigenetic control of senescence. | Peters G et al |
| 15112341 | 2004 | Association of low p16INK4a and p15INK4b mRNAs expression with their CpG islands methylation with human hepatocellular carcinogenesis. | Qin Y et al |
| 14976051 | 2004 | The interferon regulatory factor ICSBP/IRF-8 in combination with PU.1 up-regulates expression of tumor suppressor p15(Ink4b) in murine myeloid cells. | Schmidt M et al |
| 11593429 | 2001 | Deregulated c-Myb expression in murine myeloid leukemias prevents the up-regulation of p15(INK4b) normally associated with differentiation. | Schmidt M et al |
| 11283614 | 2001 | TGFbeta influences Myc, Miz-1 and Smad to control the CDK inhibitor p15INK4b. | Seoane J et al |
| 18974105 | 2008 | Conspirators in a capital crime: co-deletion of p18INK4c and p16INK4a/p14ARF/p15INK4b in glioblastoma multiforme. | Solomon DA et al |
| 11283613 | 2001 | Repression of p15INK4b expression by Myc through association with Miz-1. | Staller P et al |
| 11435325 | 2001 | Expression of p15(ink4b) gene during megakaryocytic differentiation of normal and myelodysplastic hematopoietic progenitors. | Teofili L et al |
| 12750705 | 2003 | Aberrant methylation and impaired expression of the p15(INK4b) cell cycle regulatory gene in chronic myelomonocytic leukemia (CMML). | Tessema M et al |
| 9230210 | 1997 | Cloning and characterization of p10, an alternatively spliced form of p15 cyclin-dependent kinase inhibitor. | Tsubari M et al |
| 11259171 | 2001 | Three genes with different functions in transformation are regulated by c-Myb in myeloid cells. | Wolff L et al |
| 17664422 | 2007 | Cellular senescence is an important mechanism of tumor regression upon c-Myc inactivation. | Wu CH et al |
| 18185590 | 2008 | Epigenetic silencing of tumour suppressor gene p15 by its antisense RNA. | Yu W et al |
Other Information
Locus ID:
NCBI: 1030
MIM: 600431
HGNC: 1788
Ensembl: ENSG00000147883
Variants:
dbSNP: 1030
ClinVar: 1030
TCGA: ENSG00000147883
COSMIC: CDKN2B
RNA/Proteins
| Gene ID | Transcript ID | Uniprot |
|---|---|---|
| ENSG00000147883 | ENST00000276925 | P42772 |
| ENSG00000147883 | ENST00000276925 | K7PPU3 |
| ENSG00000147883 | ENST00000380142 | P42772 |
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 38265748 | 2024 | CDKN2A/B deletion in IDH-mutant astrocytomas: An evaluation by Fluorescence in-situ hybridization. | 0 |
| 38925973 | 2024 | Frequent CDKN2B/P15 and DAPK1 methylation in duodenal follicular lymphoma is related to duodenal reactive lymphoid hyperplasia. | 0 |
| 38265748 | 2024 | CDKN2A/B deletion in IDH-mutant astrocytomas: An evaluation by Fluorescence in-situ hybridization. | 0 |
| 38925973 | 2024 | Frequent CDKN2B/P15 and DAPK1 methylation in duodenal follicular lymphoma is related to duodenal reactive lymphoid hyperplasia. | 0 |
| 35973817 | 2023 | Translational significance of CDKN2A/B homozygous deletion in isocitrate dehydrogenase-mutant astrocytoma. | 10 |
| 36261081 | 2023 | Organochlorine pesticides may induce leukemia by methylation of CDKN2B and MGMT promoters and histone modifications. | 6 |
| 36400182 | 2023 | PRMT6 functionally associates with PRMT5 to promote colorectal cancer progression through epigenetically repressing the expression of CDKN2B and CCNG1. | 3 |
| 36841437 | 2023 | Loss of CDKN2A/B is a Molecular Marker of High-grade Histology and is Associated with Aggressive Behavior in Acinic Cell Carcinoma. | 3 |
| 36879550 | 2023 | KIT A502_Y503 duplication mutation serves as a potential and universal target for neoantigen peptide in Chinese GIST patients. | 0 |
| 36961670 | 2023 | Joint Effects Between CDKN2B/P15 Methylation and Environmental Factors on the Susceptibility to Gastric Cancer. | 0 |
| 37261522 | 2023 | TP53 or CDKN2A/B covariation in ALK/RET/ROS1-rearranged NSCLC is associated with a high TMB, tumor immunosuppressive microenvironment and poor prognosis. | 3 |
| 37314514 | 2023 | Association of CDKN2A/B mutations, PD-1, and PD-L1 with the risk of acute lymphoblastic leukemia in children. | 0 |
| 37482988 | 2023 | Expression analysis of cytoskeleton regulator RNA and Cyclin Dependent Kinase Inhibitor 2B genes in breast cancer. | 0 |
| 37845622 | 2023 | Evaluation of rs10811661 polymorphism in CDKN2A / B in colon and gastric cancer. | 0 |
| 35973817 | 2023 | Translational significance of CDKN2A/B homozygous deletion in isocitrate dehydrogenase-mutant astrocytoma. | 10 |
Citation
Joanna Fares ; Linda Wolff ; Juraj Bies
CDKN2B (cyclin-dependent kinase inhibitor 2B (p15, inhibits CDK4))
Atlas Genet Cytogenet Oncol Haematol. 2011-01-01
Online version: http://atlasgeneticsoncology.org/gene/187/cdkn2b-(cyclin-dependent-kinase-inhibitor-2b-(p15-inhibits-cdk4))
