Identity
HGNC
LOCATION
3p11.1
IMAGE

LEGEND
Figure 1: Chromosomal location of EPHA3 (based on Ensembl Homo sapiens version 53.36o (NCBI36)).
LEGEND
Figure 2: Genomic neighbourhood of EPHA3 (based on Ensembl Homo sapiens version 53.36o (NCBI36)).
LOCUSID
ALIAS
EK4,ETK,ETK1,HEK,HEK4,TYRO4
FUSION GENES
DNA/RNA
Note
EPHA3 spans the human tile path clones CTD-2532M17, RP11-784B9 and RP11-547K2.

Figure 3: Genomic organisation of EPHA3.
Description
EPHA3 consists of 17 exons and 16 introns and spans 375kb of genomic DNA. It is the second largest of the EPH genes after EPHA6.
Transcription
Two alternatively spliced transcript variants have been described (NM_005233.5, a 5,807 nucleotide mRNA and NM_182644.2, a 2,684 nucleotide mRNA). The shorter transcript results in truncation within the extracellular domain of EphA3 and is predicted to produce a soluble protein. The 5 end of EPHA3 is associated with a CpG island, a feature common to all EPH genes. The EPHA3 promoter also lacks a TATA box and transcription initiates from multiple start sites.
Pseudogene
None identified.
Proteins
Note
The Eph receptors constitute the largest of the 20 subfamilies of human receptor tyrosine kinases. The founding member of this group was isolated originally from an erythropoietin producing hepatoma cell line.
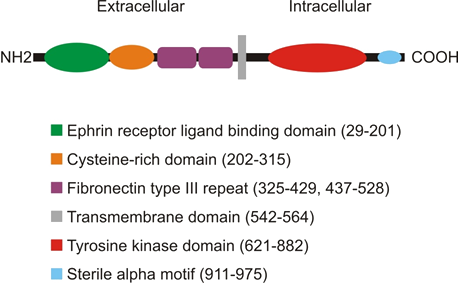

Figure 4: Domain organisation of EphA3.
Description
The EPHA3 gene encodes a 983 amino acid protein with a calculated molecular weight of 110.1kDa and an isoelectric point of 6.7302. Amino acids 1-20 constitute a signal peptide. The predicted molecular mass of the translated protein minus the signal peptide is 92.8kDa. The 521 amino acid extracellular domain contains five potential sites for N-glycosylation such that EphA3 is typically detected as a 135kDa glycoprotein. This mature isoform of EphA3 is a single-pass transmembrane receptor tyrosine kinase. Eph receptors have a conserved domain structure: At the N-terminus is a ≈ 174 amino acid ligand binding domain, followed by a ≈114 amino acid cysteine-rich domain subdivided into complement control protein (CCP, or sushi) and EGF-like domains and two membrane proximal fibronectin type III repeats. Amino acids 21-376 of the extracellular domain are rich in cysteine residues. The intracellular domain contains the tyrosine kinase domain and a sterile alpha motif. EphA3 lacks a PDZ domain interacting motif present in EphA7, EphB2, EphB3, EphB5 and EphB6. Activation of the EphA3 receptor tyrosine kinase domain is associated with two tyrosine residues in the juxtamembrane region (Y596, Y602) that are sites of autophosphorylation and interact with the kinase domain to modulate its activity.
EphA3 belongs to an evolutionarily ancient subfamily of receptor tyrosine kinases with members being present in sponges, worms and fruit flies. The expansion in the number of Eph receptor-encoding genes along with genes encoding their ligands, the ephrins (Eph receptor interacting proteins), is proposed to have contributed to the increase in complexity of the bilaterian body plan. Genes encoding EphA3 are found in the genomes of representative members of at least five of the seven classes of vertebrates including bony fish (zebrafish, pufferfish, medaka), amphibians (African clawed frog), reptiles (green anole lizard), birds (chicken) and mammals (platypus, possum, human).
Fourteen Eph receptors have been identified in vertebrates. These are subdivided into either EphA (EphA1, EphA2, EphA3, EphA4, EphA5, EphA6, EphA7, EphA8, EphA10) or EphB (EphB1, EphB2, EphB3, EphB4, EphB6) subclasses which differ primarily in the structure of their ligand binding domains. EphA receptors also exhibit greater affinity for binding GPI-linked ephrin-A ligands while EphB receptors bind transmembrane ephrin-B ligands. While interactions are somewhat promiscuous, and some cross-class binding occurs, each Eph receptor displays distinct affinity for the different ephrin ligands. The high affinity ligands for EphA3 are ephrin-A2 and ephrin-A5. EphA3 also binds ephrin-A3 and ephrin-A4 with lower affinity.
Eph-ephrin binding involves contact between cells. Upon binding, receptor-ligand dimers form heterotetramers, which further assemble into higher order signalling clusters. Several moieties in the EphA3 receptor extracellular region mediate ephrin binding. A high-affinity binding site in the N-terminal ephrin binding domain mediates intercellular Eph-ephrin interaction. Structural studies show the interaction between ephrin-A5 and EphA3 is slightly tilted relative to its binding to EphA2, resulting in a greater interaction surface . Mutation studies show two additional lower-affinity ephrin-binding sites, one in the ephrin-binding domain and the other in the cysteine-rich region, are involved in clustering of the EphA3-ephrin-A5 complex. Receptor clustering is further facilitated by receptor-receptor interactions within the ligand-binding domain and the adjacent cysteine-rich domain , which can also lead to heterologous clustering with different Eph subtypes .
Following ephrin-A5-mediated EphA3 receptor clustering, intracellular signalling by EphA3 receptors is initiated by autophosphorylation of three defined tyrosine residues, two in the highly conserved juxtamembrane region and the third in the activation loop of the kinase domain (Y779). Rapid reorganisation of the actin and myosin cytoskeleton follows, leading to retraction of cellular protrusions, membrane blebbing and cell detachment, following association of the adaptor protein CrkII with tyrosine phosphorylated EphA3 and activation of RhoA signalling.
Such Eph-ephrin interaction triggers bidirectional signalling, that is signalling events within both Eph- and ephrin-bearing cells, an unusual phenomenon for receptor tyrosine kinases, most of which interact with soluble ligands. Subsequently, depending on the cellular context (including the identity of the interacting Eph-ephrin receptor-ligand pairs, their relative levels on interacting cells, the presence of additional Ephs and ephrins and their alternative isoforms, and the net effect of interaction with additional signalling pathways) this either results in repulsion or promotes adhesion of the interacting cells.
Cellular repulsion and the termination of Eph-ephrin signalling require disruption of the receptor-ligand complex. This is brought about either by enzymatic cleavage of the tethered ephrin ligand or by trans endocytosis of Eph-ephrin complexes. EphA3-ephrin-A receptor-ligand complexes are disrupted following receptor-ligand binding when ADAM10 (a disintegrin and metalloprotease 10), cleaves ephrin cleaves ephrin, to allow cells to separate. ADAM10 association via its substrate recognition motif, and cleavage of ephrin, are dependent on ephrin/EphA3 binding and on EphA3 kinase activation. The post-cleavage ephrin-A5-EphA3 complex is then endocytosed by the EphA3-expressing cell.
While cellular repulsion is often the outcome of Eph-ephrin interaction, in some circumstances adhesion may persist, particularly when there is low Eph kinase activity. For example, ADAM10 has been observed not to cleave ephrin-A5 following EphA3-ephrin-A5 interaction involving LK63 cells in which high intracellular protein tyrosine phosphatase activity also appears to counter ephrin-A5 stimulated phosphorylation of EphA3, holding the receptor in an inactive, unphosphorylated state. The phosphatase PTP1B is known to directly regulate EphA3 activity, and its overexpression inhibits receptor endocytosis at cell-cell contacts . Another mechanism that may favour stable cell-cell adhesion involves truncated Eph receptor isoforms acting in a dominant negative manner. While activation of full length EphA7 by ephrin-A5 results in cellular repulsion, ephrin-A5-induced phosphorylation of EphA7 is inhibited by expression of two EphA7 splice variants with truncated kinase domains, which act in a dominant negative manner, and adhesion results. A splice variant of EPHA3 also has been reported, truncated before the transmembrane domain, and predicted to give rise to a soluble isoform of EphA3, the function of which has not been established. In addition, a number of EphA3 mutations have been described in various cancers (see below), at least some of which inhibit activity and cell-surface localisation of the receptor , and can act as dominant negative mutants to inhibit activity of Wt receptor .
While important details of EphA3 signalling have been determined, more complete understanding of EphA3 activity will require knowledge of the full complement of EphA3 interacting proteins. Substrates that are targets for the tyrosine kinase activity of EphA3 have yet to be defined and potential mediators or modulators of EphA3 signalling output such as Src family kinases, additional phosphotyrosine binding adaptors, SAM domain interacting factors, interaction with other receptor kinases and crosstalk with other signalling pathways, and the regulatory role of phosphatases all remain to be explored. Based on the range of interacting proteins identified for other Eph receptors (some common to more than one Eph, others apparently unique to individual Ephs) additional effectors of EphA3 signalling output are likely.
EphA3 belongs to an evolutionarily ancient subfamily of receptor tyrosine kinases with members being present in sponges, worms and fruit flies. The expansion in the number of Eph receptor-encoding genes along with genes encoding their ligands, the ephrins (Eph receptor interacting proteins), is proposed to have contributed to the increase in complexity of the bilaterian body plan. Genes encoding EphA3 are found in the genomes of representative members of at least five of the seven classes of vertebrates including bony fish (zebrafish, pufferfish, medaka), amphibians (African clawed frog), reptiles (green anole lizard), birds (chicken) and mammals (platypus, possum, human).
Fourteen Eph receptors have been identified in vertebrates. These are subdivided into either EphA (EphA1, EphA2, EphA3, EphA4, EphA5, EphA6, EphA7, EphA8, EphA10) or EphB (EphB1, EphB2, EphB3, EphB4, EphB6) subclasses which differ primarily in the structure of their ligand binding domains. EphA receptors also exhibit greater affinity for binding GPI-linked ephrin-A ligands while EphB receptors bind transmembrane ephrin-B ligands. While interactions are somewhat promiscuous, and some cross-class binding occurs, each Eph receptor displays distinct affinity for the different ephrin ligands. The high affinity ligands for EphA3 are ephrin-A2 and ephrin-A5. EphA3 also binds ephrin-A3 and ephrin-A4 with lower affinity.
Eph-ephrin binding involves contact between cells. Upon binding, receptor-ligand dimers form heterotetramers, which further assemble into higher order signalling clusters. Several moieties in the EphA3 receptor extracellular region mediate ephrin binding. A high-affinity binding site in the N-terminal ephrin binding domain mediates intercellular Eph-ephrin interaction. Structural studies show the interaction between ephrin-A5 and EphA3 is slightly tilted relative to its binding to EphA2, resulting in a greater interaction surface . Mutation studies show two additional lower-affinity ephrin-binding sites, one in the ephrin-binding domain and the other in the cysteine-rich region, are involved in clustering of the EphA3-ephrin-A5 complex. Receptor clustering is further facilitated by receptor-receptor interactions within the ligand-binding domain and the adjacent cysteine-rich domain , which can also lead to heterologous clustering with different Eph subtypes .
Following ephrin-A5-mediated EphA3 receptor clustering, intracellular signalling by EphA3 receptors is initiated by autophosphorylation of three defined tyrosine residues, two in the highly conserved juxtamembrane region and the third in the activation loop of the kinase domain (Y779). Rapid reorganisation of the actin and myosin cytoskeleton follows, leading to retraction of cellular protrusions, membrane blebbing and cell detachment, following association of the adaptor protein CrkII with tyrosine phosphorylated EphA3 and activation of RhoA signalling.
Such Eph-ephrin interaction triggers bidirectional signalling, that is signalling events within both Eph- and ephrin-bearing cells, an unusual phenomenon for receptor tyrosine kinases, most of which interact with soluble ligands. Subsequently, depending on the cellular context (including the identity of the interacting Eph-ephrin receptor-ligand pairs, their relative levels on interacting cells, the presence of additional Ephs and ephrins and their alternative isoforms, and the net effect of interaction with additional signalling pathways) this either results in repulsion or promotes adhesion of the interacting cells.
Cellular repulsion and the termination of Eph-ephrin signalling require disruption of the receptor-ligand complex. This is brought about either by enzymatic cleavage of the tethered ephrin ligand or by trans endocytosis of Eph-ephrin complexes. EphA3-ephrin-A receptor-ligand complexes are disrupted following receptor-ligand binding when ADAM10 (a disintegrin and metalloprotease 10), cleaves ephrin cleaves ephrin, to allow cells to separate. ADAM10 association via its substrate recognition motif, and cleavage of ephrin, are dependent on ephrin/EphA3 binding and on EphA3 kinase activation. The post-cleavage ephrin-A5-EphA3 complex is then endocytosed by the EphA3-expressing cell.
While cellular repulsion is often the outcome of Eph-ephrin interaction, in some circumstances adhesion may persist, particularly when there is low Eph kinase activity. For example, ADAM10 has been observed not to cleave ephrin-A5 following EphA3-ephrin-A5 interaction involving LK63 cells in which high intracellular protein tyrosine phosphatase activity also appears to counter ephrin-A5 stimulated phosphorylation of EphA3, holding the receptor in an inactive, unphosphorylated state. The phosphatase PTP1B is known to directly regulate EphA3 activity, and its overexpression inhibits receptor endocytosis at cell-cell contacts . Another mechanism that may favour stable cell-cell adhesion involves truncated Eph receptor isoforms acting in a dominant negative manner. While activation of full length EphA7 by ephrin-A5 results in cellular repulsion, ephrin-A5-induced phosphorylation of EphA7 is inhibited by expression of two EphA7 splice variants with truncated kinase domains, which act in a dominant negative manner, and adhesion results. A splice variant of EPHA3 also has been reported, truncated before the transmembrane domain, and predicted to give rise to a soluble isoform of EphA3, the function of which has not been established. In addition, a number of EphA3 mutations have been described in various cancers (see below), at least some of which inhibit activity and cell-surface localisation of the receptor , and can act as dominant negative mutants to inhibit activity of Wt receptor .
While important details of EphA3 signalling have been determined, more complete understanding of EphA3 activity will require knowledge of the full complement of EphA3 interacting proteins. Substrates that are targets for the tyrosine kinase activity of EphA3 have yet to be defined and potential mediators or modulators of EphA3 signalling output such as Src family kinases, additional phosphotyrosine binding adaptors, SAM domain interacting factors, interaction with other receptor kinases and crosstalk with other signalling pathways, and the regulatory role of phosphatases all remain to be explored. Based on the range of interacting proteins identified for other Eph receptors (some common to more than one Eph, others apparently unique to individual Ephs) additional effectors of EphA3 signalling output are likely.
Expression
EphA3 was first identified as an antigen expressed at high levels (10,000-20,000 copies per cell) on the surface of the LK63 pre-B cell acute lymphoblastic leukaemia cell line. It also was found to be expressed by JM, HSB-2 and MOLT-4 T-cell leukaemic cell lines, in CD28-stimulated Jurkat cells, and in 16 of 42 cases of primary T-cell lymphoma (but not normal peripheral T lymphocytes nor in any subset of thymus-derived developing T cells). It is also present in patients with hematologic malignancies, including AML, CML, MDS, MPN and Multiple Myeloma, and in various solid tumours (see below)
EphA3 expression has been shown to be most abundant during vertebrate development, where it is highly regulated both temporally and spatially. Prominent EphA3 expression occurs in the neural system, including the retinal ganglion cells of the embryonic retina in a graded distribution from anterior/nasal (lowest) to posterior/temporal (highest); the cerebrum, thalamus, striatum, olfactory bulb, anterior commissure, and corpus callosum of the forebrain; and the medial motor column ventral motor neurons of the spinal cord; and extraneurally by mesodermally-derived tissues including the paraxial musculature, tongue musculature, submucosa of the soft palate, capsule of the submandibular gland, cortical rim of bone, thymic septae, media of the pharynx, trachea, great vessels, small intestine and portal vein, cardiac valves, and the renal medulla. In adult tissues EphA3 expression is more restricted and detected at significantly lower levels than during early development. However it is expressed on mesenchymal stromal progenitor cells (MSCs) during neovascularisation of the regenerating endometrium, with its expression regulated by hypoxia. Similarly EphA3 is expressed on MSCs recruited from the bone-marrow and contributing to the vasculature and supporting stromal tissue of various solid cancers . It is also over-expressed on progenitor cells in gliomas. In these instances EphA3 in tumours is largely inactive, and activation with an agonistic antibody inhibits tumour growth.
Loss of EphA3 expression is also reported in cancer. EPHA3 gene copy number and/or expression levels were decreased in lung cancers (157 of 371 primary lung adenocarcinomas), and also in esophageal squamous cell carcinoma (ESCC). Silencing of EPHA3 expression by DNA hypermethylation occurs in leukaemia , and in colorectal tumours carrying a BRAF mutation (V600E). Somatic mutations identified in the 3 untranslated region of EPHA3 may also disrupt miRNA target sites, thereby altering its expression .
EphA3 expression has been shown to be most abundant during vertebrate development, where it is highly regulated both temporally and spatially. Prominent EphA3 expression occurs in the neural system, including the retinal ganglion cells of the embryonic retina in a graded distribution from anterior/nasal (lowest) to posterior/temporal (highest); the cerebrum, thalamus, striatum, olfactory bulb, anterior commissure, and corpus callosum of the forebrain; and the medial motor column ventral motor neurons of the spinal cord; and extraneurally by mesodermally-derived tissues including the paraxial musculature, tongue musculature, submucosa of the soft palate, capsule of the submandibular gland, cortical rim of bone, thymic septae, media of the pharynx, trachea, great vessels, small intestine and portal vein, cardiac valves, and the renal medulla. In adult tissues EphA3 expression is more restricted and detected at significantly lower levels than during early development. However it is expressed on mesenchymal stromal progenitor cells (MSCs) during neovascularisation of the regenerating endometrium, with its expression regulated by hypoxia. Similarly EphA3 is expressed on MSCs recruited from the bone-marrow and contributing to the vasculature and supporting stromal tissue of various solid cancers . It is also over-expressed on progenitor cells in gliomas. In these instances EphA3 in tumours is largely inactive, and activation with an agonistic antibody inhibits tumour growth.
Loss of EphA3 expression is also reported in cancer. EPHA3 gene copy number and/or expression levels were decreased in lung cancers (157 of 371 primary lung adenocarcinomas), and also in esophageal squamous cell carcinoma (ESCC). Silencing of EPHA3 expression by DNA hypermethylation occurs in leukaemia , and in colorectal tumours carrying a BRAF mutation (V600E). Somatic mutations identified in the 3 untranslated region of EPHA3 may also disrupt miRNA target sites, thereby altering its expression .
Localisation
Isoform 1: Cell membrane; single-pass type I membrane protein.
Isoform 2: Secreted.
Isoform 2: Secreted.
Function
Eph receptors modulate cell shape and movement through reorganisation of the cytoskeleton and changes in cell-cell and cell-substrate adhesion, and are involved in many cellular migration, sorting (tissue patterning) and guidance events, most often during development, and in particular involving the nervous system. There is evidence too that Eph receptor signalling influences cell proliferation and cell-fate determination and growing recognition that Eph receptors function in adult tissue homeostasis.
EphA3 is thought to play a role in retinotectal mapping, the tightly patterned projection of retinal ganglion cell axons from the retina to the optic tectum (or superior colliculus in mammals). In chicks, posterior retinal ganglion axons expressing highest levels of EphA3 project to the anterior tectum where the graded expression of ephrin-A2 and ephrin-A5 is lowest and are excluded from projecting more posteriorly where ephrin-A2/A5 expression is highest. More direct evidence of non-redundant function for EphA3 has come from phenotypic analysis of EphA3 knockout mice. Approximately 70-75% of EphA3 null mice die within 48 hours of birth with post-mortem evidence of pulmonary oedema secondary to cardiac failure. These mice exhibit hypoplastic atrioventricular endocardial cushions and subsequent atrioventricular valve and atrial membranous septal defects, with endocardial cushion explants from these mice giving rise to fewer migrating cells arising from epithelial to mesenchymal transformation. Expression of EphA3 in the spinal cord appears to be redundant as axial muscle targeting by medial motor column motor axons and the organisation of the motor neuron columns is not altered. EphA4 is the only other EphA receptor also expressed by developing spinal cord motor neurons and in mice lacking EphA3 and EphA4 these receptors together repel axial motor axons from neighbouring ephrin-A-expressing sensory axons, inhibiting intermingling of motor and sensory axons and preventing mis-projection of motor axons into the dorsal root ganglia. In contrast to the chick, EphA3 is not expressed by mouse retinal ganglion cells. Instead the closely related receptors EphA5 and EphA6 (see homology below) are expressed in a low nasal to high temporal gradient. However, if EphA3 is ectopically expressed in retinal ganglion cells in mice these axons project to more rostral positions in the superior colliculus.
A function for soluble EphA3 has not been reported although potentially this isoform might play a role in promoting cell adhesion (see above) or act as a tumour suppressor protein (see below).
EphA3 is thought to play a role in retinotectal mapping, the tightly patterned projection of retinal ganglion cell axons from the retina to the optic tectum (or superior colliculus in mammals). In chicks, posterior retinal ganglion axons expressing highest levels of EphA3 project to the anterior tectum where the graded expression of ephrin-A2 and ephrin-A5 is lowest and are excluded from projecting more posteriorly where ephrin-A2/A5 expression is highest. More direct evidence of non-redundant function for EphA3 has come from phenotypic analysis of EphA3 knockout mice. Approximately 70-75% of EphA3 null mice die within 48 hours of birth with post-mortem evidence of pulmonary oedema secondary to cardiac failure. These mice exhibit hypoplastic atrioventricular endocardial cushions and subsequent atrioventricular valve and atrial membranous septal defects, with endocardial cushion explants from these mice giving rise to fewer migrating cells arising from epithelial to mesenchymal transformation. Expression of EphA3 in the spinal cord appears to be redundant as axial muscle targeting by medial motor column motor axons and the organisation of the motor neuron columns is not altered. EphA4 is the only other EphA receptor also expressed by developing spinal cord motor neurons and in mice lacking EphA3 and EphA4 these receptors together repel axial motor axons from neighbouring ephrin-A-expressing sensory axons, inhibiting intermingling of motor and sensory axons and preventing mis-projection of motor axons into the dorsal root ganglia. In contrast to the chick, EphA3 is not expressed by mouse retinal ganglion cells. Instead the closely related receptors EphA5 and EphA6 (see homology below) are expressed in a low nasal to high temporal gradient. However, if EphA3 is ectopically expressed in retinal ganglion cells in mice these axons project to more rostral positions in the superior colliculus.
A function for soluble EphA3 has not been reported although potentially this isoform might play a role in promoting cell adhesion (see above) or act as a tumour suppressor protein (see below).
Homology
Phylogenetic tree for the Eph receptors. Amino acid sequences used for this compilation were EphA1 (NP_005223), EphA2 (NM_004431), EphA3 (NP_005224), EphA4 (NP_004429), EphA5 (NM_004439), EphA6 (ENSP00000374323), EphA7 (NP_004431), EphA8 (NP_065387), EphA10 (NP_001092909), EphB1 (NP_004432), EphB2 (NP_004433), EphB3 (NP_004434), EphB4 (NP_004435) and EphB6 (NP_004436).

Mutations
Note
.
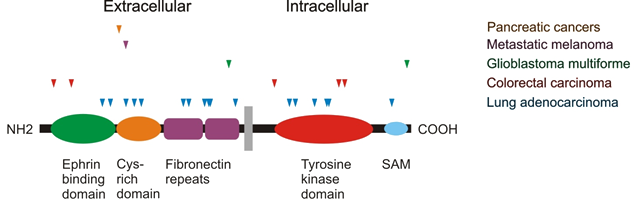
Figure 6: Sites of somatic mutations in EphA3 identified in lung adenocarcinoma colorectal carcinoma, glioblastoma multiforme, metastatic melanoma and pancreatic cancers (PDAC and AVC).
Germinal
To date no germinal mutations in EPHA3 have been associated with disease.
Somatic
Somatic mutations in EPHA3 have been frequently detected, including in lung adenocarcinoma (T166N, G187R, S229Y, W250R, M269I, N379K, T393K, A435S, D446Y, S449F, G518L, T660K, D678E, R728L, K761N, G766E, T933M), colorectal carcinoma (T37K, N85S, I621L, S792P, D806N), glioblastoma multiforme (K500N, A971P) metastatic melanoma (G228R) and pancreatic cancer (K207N). Two mutations are reported in haematological malignancies: R897M in mantle cell lymphoma and a truncating E461X mutation in a t(4;14) myeloma (with FGFR3/ WHSC1 (MMSET) translocation). According to the COSMIC catalogue of somatic mutations in cancer (http://cancer.sanger.ac.uk) there are over 300 reported mutations, which are distributed over all regions encoding functional domains, most prominently in the extracellular domains involved in ligand-receptor and receptor-receptor interactions, and in the intracellular kinase domain. Several mutations have been confirmed to inhibit receptor activity and cell surface expression..
Implicated in
Entity name
Prostate cancer
Note
EPHA3 was among the genes whose expression was upregulated during androgen-independent progression in an LNCaP in vitro cell model of prostate cancer. Subsequently, EphA3 was found to correlate with proliferation and survival of prostate cancer cells and tumour growth in mice, and was upregulated in stromal cells at sites of bone metastasis.
Entity name
Melanoma
Note
A melanoma patient with an especially favourable evolution of disease, associated with a very strong and sustained anti-tumour cytotoxic T lymphocyte response, was found to have a lytic CD4 clone that recognised an EphA3 antigen presented by the HLA class II molecule HLA- DRB1*1101. 94% (75 of 80) of melanomas examined expressed EphA3 in contrast to normal melanocytes which do not express detectable EphA3.
Entity name
Lung cancer, Sarcoma, and Renal cell carcinoma
Note
44% (11 of 25) of small cell lung cancer, 24% (10 of 41) of non-small cell lung cancer, 58% (17 of 29) of sarcomas, and 31% (12 of 38) of renal cell carcinomas expressed EphA3 at levels significantly higher than the corresponding normal tissues.
Entity name
Liver, gastric, and colorectal cancer
Note
High EphA3 expression associated with high invasive capacity and poor overall survival in hepatocellular carcinoma , and with angiogenesis and poor prognosis in gastric cancer. In colorectal cancer, high expression correlates with stem cell marker expression, and with tumour size and grade, infiltration and metastasis.
Entity name
Hematological tumours
Note
Increased expression in 50% of patients with myelodysplastic syndrome, acute myeloid leukaemia or chronic myeloid leukaemia (CML), most prominent on a leukaemia stem cell immunophenotype. Overexpression in a high proportion of the other chronic myeloproliferative diseases was also observed.
Entity name
Glioma
Note
40% glioma specimens over-expressed EphA3, particularly in mesenchymal subtype. Expression on cancer initiating/stem cell type. Radio-labelled antibody targeting in mouse model inhibited growth.
Entity name
Other solid tumours: bladder, brain, breast, colon, kidney, liver, lung, melanoma, prostate
Note
Over-expressed in high proportion of tumour vasculature and stromal tissues, even when not in the tumour bulk. Expressed on mesenchymal stromal cells recruited from bone marrow. Targeting with activating antibody inhibited tumour growth in mouse models.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 25485970 | 2014 | The molecular signature of the stroma response in prostate cancer-induced osteoblastic bone metastasis highlights expansion of hematopoietic and prostate epithelial stem cell niches. | Özdemir BC et al |
| 16785212 | 2006 | Approximate likelihood-ratio test for branches: A fast, accurate, and powerful alternative. | Anisimova M et al |
| 17440062 | 2007 | Novel somatic and germline mutations in cancer candidate genes in glioblastoma, melanoma, and pancreatic carcinoma. | Balakrishnan A et al |
| 12738854 | 2003 | Mutational analysis of the tyrosine kinome in colorectal cancers. | Bardelli A et al |
| 24145436 | 2013 | Landscape of somatic mutations and clonal evolution in mantle cell lymphoma. | Beà S et al |
| 11741094 | 2001 | Signals from Eph and ephrin proteins: a developmental tool kit. | Boyd AW et al |
| 1737782 | 1992 | Isolation and characterization of a novel receptor-type protein tyrosine kinase (hek) from a human pre-B cell line. | Boyd AW et al |
| 12370823 | 2002 | Soluble Eph A receptors inhibit tumor angiogenesis and progression in vivo. | Brantley DM et al |
| 10929715 | 2000 | Topographic mapping from the retina to the midbrain is controlled by relative but not absolute levels of EphA receptor signaling. | Brown A et al |
| 16491080 | 2006 | Silencing of EphA3 through a cis interaction with ephrinA5. | Carvalho RF et al |
| 10742046 | 2000 | Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. | Castresana J et al |
| 18241037 | 2008 | Genomic profiling of 766 cancer-related genes in archived esophageal normal and carcinoma tissues. | Chen J et al |
| 14670182 | 2003 | Inhibition of VEGF-dependent multistage carcinogenesis by soluble EphA receptors. | Cheng N et al |
| 17032440 | 2006 | TreeDyn: towards dynamic graphics and annotations for analyses of trees. | Chevenet F et al |
| 10987298 | 2000 | Identification of a tumor-specific shared antigen derived from an Eph receptor and presented to CD4 T cells on HLA class II molecules. | Chiari R et al |
| 20838624 | 2010 | Mutational profiling of kinases in human tumours of pancreatic origin identifies candidate cancer genes in ductal and ampulla of vater carcinomas. | Corbo V et al |
| 16140923 | 2005 | Somatic mutations of the protein kinase gene family in human lung cancer. | Davies H et al |
| 18547520 | 2008 | Autoregulation by the juxtamembrane region of the human ephrin receptor tyrosine kinase A3 (EphA3). | Davis TL et al |
| 15901737 | 2005 | Three distinct molecular surfaces in ephrin-A5 are essential for a functional interaction with EphA3. | Day B et al |
| 25144626 | 2014 | Eph receptors as therapeutic targets in glioblastoma. | Day BW et al |
| 18424797 | 2008 | Phylogeny.fr: robust phylogenetic analysis for the non-specialist. | Dereeper A et al |
| 18948947 | 2008 | Somatic mutations affect key pathways in lung adenocarcinoma. | Ding L et al |
| 10498621 | 1999 | Cloning and characterization of EphA3 (Hek) gene promoter: DNA methylation regulates expression in hematopoietic tumor cells. | Dottori M et al |
| 12100883 | 2002 | Eph family functions from an evolutionary perspective. | Drescher U et al |
| 15034147 | 2004 | MUSCLE: multiple sequence alignment with high accuracy and high throughput. | Edgar RC et al |
| 15014130 | 2004 | Loss-of-function analysis of EphA receptors in retinotectal mapping. | Feldheim DA et al |
| 25993310 | 2015 | Distinctive Structure of the EphA3/Ephrin-A5 Complex Reveals a Dual Mode of Eph Receptor Interaction for Ephrin-A5. | Forse GJ et al |
| 18403711 | 2008 | Segregation of axial motor and sensory pathways via heterotypic trans-axonal signaling. | Gallarda BW et al |
| 17344846 | 2007 | Patterns of somatic mutation in human cancer genomes. | Greenman C et al |
| 14530136 | 2003 | A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. | Guindon S et al |
| 14726470 | 2004 | Differential gene expression of Eph receptors and ephrins in benign human tissues and cancers. | Hafner C et al |
| 10958785 | 2000 | Regulated cleavage of a contact-mediated axon repellent. | Hattori M et al |
| 20505120 | 2010 | Architecture of Eph receptor clusters. | Himanen JP et al |
| 20027224 | 2009 | Analysis of the association between CIMP and BRAF in colorectal cancer by DNA methylation profiling. | Hinoue T et al |
| 2825356 | 1987 | A novel putative tyrosine kinase receptor encoded by the eph gene. | Hirai H et al |
| 9707552 | 1998 | PDZ-domain-mediated interaction of the Eph-related receptor tyrosine kinase EphB3 and the ras-binding protein AF6 depends on the kinase activity of the receptor. | Hock B et al |
| 22144690 | 2011 | Eph receptor function is modulated by heterooligomerization of A and B type Eph receptors. | Janes PW et al |
| 16239146 | 2005 | Adam meets Eph: an ADAM substrate recognition module acts as a molecular switch for ephrin cleavage in trans. | Janes PW et al |
| 25391995 | 2014 | EphA3 biology and cancer. | Janes PW et al |
| 22780212 | 2012 | EPHA3 as a novel therapeutic target in the hematological malignancies. | Keane N et al |
| 8812059 | 1996 | Expression of the Tyro4/Mek4/Cek4 gene specifically marks a subset of embryonic motor neurons and their muscle targets. | Kilpatrick TJ et al |
| 15892098 | 2005 | Expression profiles of EphA3 at both the RNA and protein level in the developing mammalian forebrain. | Kudo C et al |
| 9685371 | 1998 | Distinct subdomains of the EphA3 receptor mediate ligand binding and receptor dimerization. | Lackmann M et al |
| 11870224 | 2002 | Ephrin-A5 induces rounding, blebbing and de-adhesion of EphA3-expressing 293T and melanoma cells by CrkII and Rho-mediated signalling. | Lawrenson ID et al |
| 22242939 | 2012 | Cancer somatic mutations disrupt functions of the EphA3 receptor tyrosine kinase through multiple mechanisms. | Lisabeth EM et al |
| 23970317 | 2013 | High levels of EphA3 expression are associated with high invasive capacity and poor overall survival in hepatocellular carcinoma. | Lu CY et al |
| 21135139 | 2010 | PTP1B regulates Eph receptor function and trafficking. | Nievergall E et al |
| 18394988 | 2008 | Eph-ephrin bidirectional signaling in physiology and disease. | Pasquale EB et al |
| 15469835 | 2004 | Diverse roles of eph receptors and ephrins in the regulation of cell migration and tissue assembly. | Poliakov A et al |
| 11114734 | 2000 | The protein tyrosine kinase family of the human genome. | Robinson DR et al |
| 20228801 | 2010 | An extracellular steric seeding mechanism for Eph-ephrin signaling platform assembly. | Seiradake E et al |
| 17977648 | 2008 | Genome-wide expression profiling reveals transcriptomic variation and perturbed gene networks in androgen-dependent and androgen-independent prostate cancer cells. | Singh AP et al |
| 16959974 | 2006 | The consensus coding sequences of human breast and colorectal cancers. | Sjöblom T et al |
| 14660665 | 2004 | Dissecting the EphA3/Ephrin-A5 interactions using a novel functional mutagenesis screen. | Smith FM et al |
| 14697337 | 2004 | EphA3 is induced by CD28 and IGF-1 and regulates cell adhesion. | Smith LM et al |
| 17046737 | 2007 | A critical role for the EphA3 receptor tyrosine kinase in heart development. | Stephen LJ et al |
| 25420155 | 2014 | Hypoxia-controlled EphA3 marks a human endometrium-derived multipotent mesenchymal stromal cell that supports vascular growth. | To C et al |
| 14585969 | 2003 | EphA3 null mutants do not demonstrate motor axon guidance defects. | Vaidya A et al |
| 25125683 | 2014 | Targeting EphA3 inhibits cancer growth by disrupting the tumor stromal microenvironment. | Vail ME et al |
| 16061656 | 2005 | Concurrent binding of anti-EphA3 antibody and ephrin-A5 amplifies EphA3 signaling and downstream responses: potential as EphA3-specific tumor-targeting reagents. | Vearing C et al |
| 22573403 | 2012 | Intraclonal heterogeneity and distinct molecular mechanisms characterize the development of t(4;14) and t(11;14) myeloma. | Walker BA et al |
| 1311845 | 1992 | Molecular cloning of HEK, the gene encoding a receptor tyrosine kinase expressed by human lymphoid tumor cell lines. | Wicks IP et al |
| 18385452 | 2008 | Elevated protein tyrosine phosphatase activity provokes Eph/ephrin-facilitated adhesion of pre-B leukemia cells. | Wimmer-Kleikamp SH et al |
| 16941478 | 2006 | Somatic mutations of GUCY2F, EPHA3, and NTRK3 in human cancers. | Wood LD et al |
| 25231727 | 2014 | EphA3, induced by PC-1/PrLZ, contributes to the malignant progression of prostate cancer. | Wu R et al |
| 22350700 | 2012 | Aberrant expression of EphA3 in gastric carcinoma: correlation with tumor angiogenesis and survival. | Xi HQ et al |
| 21415057 | 2011 | Clinicopathological significance and prognostic value of EphA3 and CD133 expression in colorectal carcinoma. | Xi HQ et al |
| 22829656 | 2012 | Effects of cancer-associated EPHA3 mutations on lung cancer. | Zhuang G et al |
| 23091610 | 2012 | Integrative analysis of somatic mutations altering microRNA targeting in cancer genomes. | Ziebarth JD et al |
Other Information
Locus ID:
NCBI: 2042
MIM: 179611
HGNC: 3387
Ensembl: ENSG00000044524
Variants:
dbSNP: 2042
ClinVar: 2042
TCGA: ENSG00000044524
COSMIC: EPHA3
RNA/Proteins
| Gene ID | Transcript ID | Uniprot |
|---|---|---|
| ENSG00000044524 | ENST00000336596 | P29320 |
| ENSG00000044524 | ENST00000336596 | A0A140VJJ0 |
| ENSG00000044524 | ENST00000452448 | P29320 |
| ENSG00000044524 | ENST00000494014 | C9JXA2 |
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 37434266 | 2023 | The Ephrin tyrosine kinase a3 (EphA3) is a novel mediator of RAGE-prompted motility of breast cancer cells. | 2 |
| 37434266 | 2023 | The Ephrin tyrosine kinase a3 (EphA3) is a novel mediator of RAGE-prompted motility of breast cancer cells. | 2 |
| 33417456 | 2021 | Intranasal Delivery of Temozolomide-Conjugated Gold Nanoparticles Functionalized with Anti-EphA3 for Glioblastoma Targeting. | 14 |
| 33919657 | 2021 | EPHA3 Contributes to Epigenetic Suppression of PTEN in Radioresistant Head and Neck Cancer. | 5 |
| 33417456 | 2021 | Intranasal Delivery of Temozolomide-Conjugated Gold Nanoparticles Functionalized with Anti-EphA3 for Glioblastoma Targeting. | 14 |
| 33919657 | 2021 | EPHA3 Contributes to Epigenetic Suppression of PTEN in Radioresistant Head and Neck Cancer. | 5 |
| 31433759 | 2020 | Sam68 Promotes the Progression of Human Breast Cancer through inducing Activation of EphA3. | 8 |
| 32621117 | 2020 | Critical role of EphA3 in cancer and current state of EphA3 drug therapeutics. | 13 |
| 32633364 | 2020 | EphA3 promotes the proliferation of NPC cells through negatively regulating the ability of FOG2. | 1 |
| 31433759 | 2020 | Sam68 Promotes the Progression of Human Breast Cancer through inducing Activation of EphA3. | 8 |
| 32621117 | 2020 | Critical role of EphA3 in cancer and current state of EphA3 drug therapeutics. | 13 |
| 32633364 | 2020 | EphA3 promotes the proliferation of NPC cells through negatively regulating the ability of FOG2. | 1 |
| 30483759 | 2019 | EphA3 inhibits migration and invasion of esophageal cancer cells by activating the mesenchymal‑epithelial transition process. | 9 |
| 30528229 | 2019 | EphA3 is up-regulated by epidermal growth factor and promotes formation of glioblastoma cell aggregates. | 1 |
| 30560328 | 2019 | EphA3 Downregulation by Hypermethylation Associated with Lymph Node Metastasis and TNM Stage in Colorectal Cancer. | 7 |
Citation
Peter W. Janes
EPHA3 (EPH receptor A3)
Atlas Genet Cytogenet Oncol Haematol. 2015-07-01
Online version: http://atlasgeneticsoncology.org/gene/40463/epha3-(eph-receptor-a3)
Historical Card
2009-04-01 EPHA3 (EPH receptor A3) by Brett Stringer,Bryan Day,Jennifer McCarron,Martin Lackmann,Andrew Boyd Affiliation
