AXIN1 (axin 1)
2011-05-01 Nives Pecina-Slaus , Tamara Nikuseva Martic , Tomislav Kokotovic AffiliationDepartment of Biology, Laboratory for Neurooncology, Croatian Institute for Brain Research, Medical School University of Zagreb, Salata 12, Zagreb, Croatia
DNA/RNA
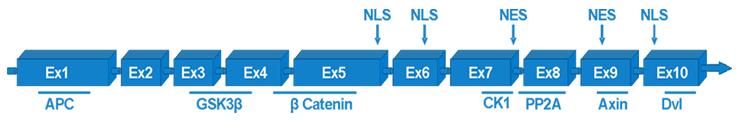
Figure 1. Genomic structure of Axin 1. Axin 1 is composed of 10 exons and they encode isoform a, while in isoform b exon 8 is spliced out.
Description
Axin 1 consists of 11 exons (isoform a). Full gene transcript product length is 3675 bp. Isoform b lacks an in-frame exon in the 3 coding region and is shorter with sequence length of 3567 bp (Salahshor and Woodgett, 2005) (Figure 1).
Transcription
There are two transcript variants. Variant 1 (encoding for isoform a) represents the longer transcript (NM 003502.3). Variant 2 (encoding for isoform b) is shorter compared to variant 1 (NM 181050.2). According to Ensembl there are six transcripts of AXIN1 of which first two are well known isoforms a and b and the remaining 4 are still in research.
Proteins
Note
Protein name: Axin 1, Axin, Axis inhibitor, Axis inhibitor protein 1.
Description
At least two isoforms of protein axin are expressed. Longer isoform has all eleven exons translated and consists of 862 aminoacids while shorter has 826 aminoacids translated from ten exons. Axin 1 protein can be recognized primarily by two domains, the N-terminal RGS domain (regulators of G-protein signaling) and the C-terminal DIX domain (dishevelled and axin) (Luo et al., 2005; Shibata et al., 2007). RGS domain is needed for APC binding while DIX domain for homodimerization and heterodimerization (Ehebauer and Arias, 2009; Noutsou et al., 2011). There is also a central region of the protein that binds GSK3beta and beta-catenin. Axin protein has nuclear localization (NLS) and nuclear export (NES) sequences as well. It is well known that axin is a scaffold protein that can shuttle between the cytoplasm and the nucleus. Nucleo-cytoplasmatic shuttling under normal circumstances suggests existence of possible "salvage pathway" that would be activated by axin translocation to the nucleus in order to reduce beta-catenin oncogenic activity by exporting nuclear beta-catenin and degrading it in the cytoplasm (Wiechens et al., 2004). Axin can also undergo posttranslational modifications. Phosphorilation by casein kinase 1 (CK1) enhances binding of GSK3beta and AXIN1. For activation of JNK pathway axin needs to be SUMOylated (Kim et al., 2008) (Figure 2).
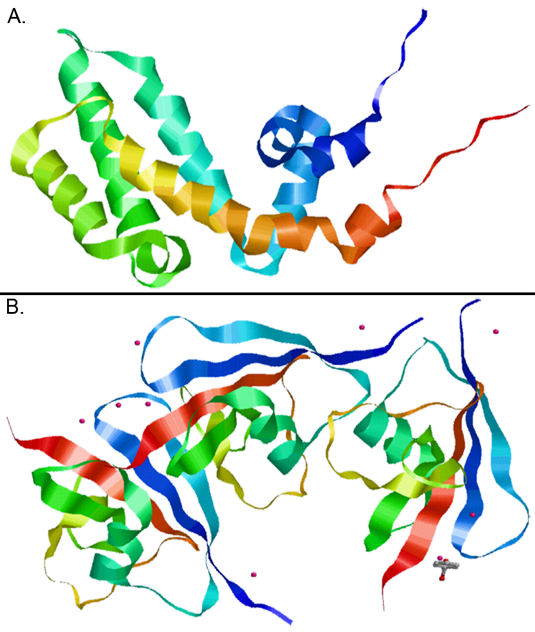
Figure 2. Two crystallized domains of the Axin 1 protein are shown: (A) RGS and (B) DIX.
Expression
Axin is expressed ubiquitously.
Localisation
Axin is predominantlly expressed in the cytoplasm, but periplasmic and nuclear localization are also observed depending on the stimulation of the cells (Cong and Varmus, 2004; Luo and Lin, 2004). In nonstimulated cells, axin colocalizes with Smad3. The subcellular location of axin is not well defined in the literature. It has been reported that physiological concentrations of axin is low in Xenopus egg cells. It has also been shown that it is located in cytoplasmic puncta in living mammalian cells. Wang et al. (2009) report that axin 1 is highly co-localized with beta-catenin in the cytoplasm of human cumulus cells and that this localization denotes intact wnt signaling. Pecina-Slaus et al. (2011) showed the subcellular location of axin in normal brain white matter and glioblastoma tissue. The majority of glioblastomas (69.04%) had axin localized in the cytoplasm. Nevertheless, 9.5% of glioblastomas samples had axin localized in the nucleus (Figure 3). Distribution of axin was reported previously by Anderson et al. (2002) in neoplastic colon. Altered nuclear expression of axin seen in colon polyps and carcinomas may be a consequence of the loss of full-length APC and the advent of nuclear beta-catenin.
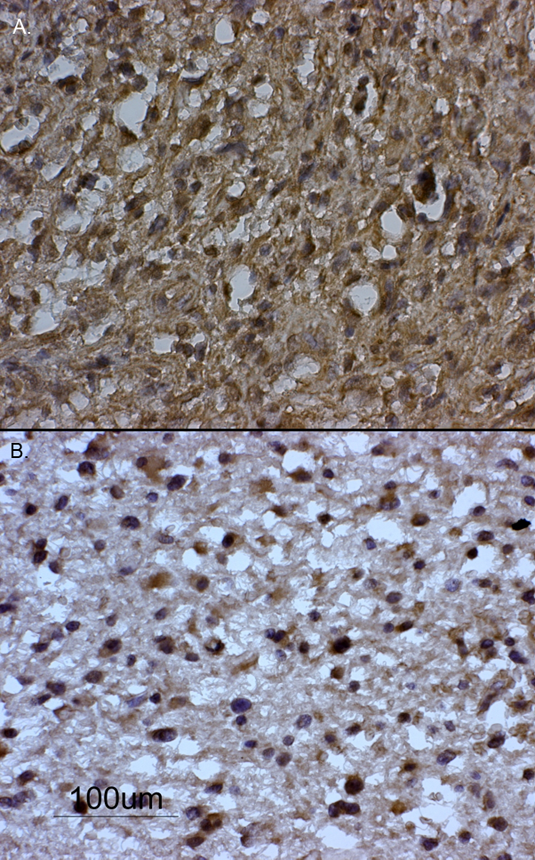
Figure 3. Glioblastoma samples immunohistochemically stained for protein expression of axin. (A) Cytoplasmic localization of axin and (B) nuclear localization of axin.
Function
Tumor suppressor protein Axin 1 is an inhibitor of the Wnt signaling pathway (Polakis, 2000; Salahshor and Woodgett, 2005). As a scaffold protein, its main role is binding multiple members of Wnt signaling and formation of the beta-catenin destruction complex. It down-regulates beta-catenin, wnt pathways main effector signaling molecule, by facilitating its phosphorylation by GSK3-beta (Hart et al., 1998). It binds directly to APC (adenomatous polyposis coli), beta-catenin, GSK3-beta and dishevelled forming a so called "beta-catenin destruction complex" in which phosphorylated beta-catenin is targeted for quick ubiquitinilation and degradation in the 26S proteosome (Yamamoto et al., 1999; Logan and Nusse, 2004). In response to wnt signaling, or under the circumstances of mutated axin or APC, beta-catenin is stabilized, accumulates in the cytoplasm and enters the nucleus, where it finds a partner, a member of the DNA binding protein family LEF/TCF. Together they stimulate the expression of target genes including c-myc, c-jun, fra-1 and cyclin D1. In developement Axin controls dorsoventral polarity axis formation (Zeng et al., 1997; Wodarz and Nusse, 1998) by two independent mechanisms: downregulation of beta-catenin, but also by activation of Wnt-independent JNK signaling activation. Axin has a role in determining cells fate upon damage, haematopoetic stem cells differentiation (Reya et al., 2003) and transforming growth factor beta signaling (Furuhashi et al., 2001). Reports indicate that beta-catenin and axin regulate critical developmental processes of normal CNS development (Pecina-Slaus, 2010).
Axin interacts with a number of proteins including: APC, Axam, Axin, beta-catenin, Ccd1, CKI, DAXX, DCAP, Diversin, Dvl, gamma-tubulin, GSK3beta, HIPK2, I-mfa, LRP5/LRP6, MDFIC, MEKK1, MEKK4, P53, PIAS, Pirh2, PP2A, Rnf11, Zbed3, Tip60, Smad3, Smad6, and Smad7 (Cliffe et al., 2003; Chen et al., 2009; Fumoto et al., 2009; Li et al., 2009; Choi et al., 2010; Kim and Jho, 2010).
Axin interacts with a number of proteins including: APC, Axam, Axin, beta-catenin, Ccd1, CKI, DAXX, DCAP, Diversin, Dvl, gamma-tubulin, GSK3beta, HIPK2, I-mfa, LRP5/LRP6, MDFIC, MEKK1, MEKK4, P53, PIAS, Pirh2, PP2A, Rnf11, Zbed3, Tip60, Smad3, Smad6, and Smad7 (Cliffe et al., 2003; Chen et al., 2009; Fumoto et al., 2009; Li et al., 2009; Choi et al., 2010; Kim and Jho, 2010).
Homology
Homologs are found in: Pan troglodytes, Canis lupus familiaris, Bos taurus, Mus musculus, Rattus norvegicus, Gallus gallus, Danio rerio.
Mutations
Note
According to HGMD there are 3 missense mutations reported for AXIN 1 in colorectal carcinoma. Nikuseva Martic et al. (2010) identified gross deletions (Loss of Heterozygosity) of AXIN 1 in 6.3% of glioblastomas, in one neuroepithelial dysembrioplastic tumor and in one medulloblastoma. In a primary hepatocellular carcinoma 13 somatic events were reported by OMIM, a 33-bp deletion in exon 3 of the AXIN1 gene, and 12 missense mutations. OMIM also reports on hypermethylation of AXIN 1 promotor region in caudal duplication anomaly.
Implicated in
Entity name
Hepatocellular carcinoma
Note
In a primary hepatocellular carcinoma (HCC), Satoh et al. (2000) found a 33-bp deletion in exon 3 of the AXIN1 gene, involving 2 glycogen synthase kinase-3-beta phosphorylation sites. In addition to this deletion they found 12 missense mutations, of which 9 occurred in codons encoding serine or threonine residues. They confirmed that all 13 mutations found in primary HCCs occurred as somatic events. Taniguchi et al. (2002) found AXIN1 mutations in seven (9.6%) HCCs. The AXIN1 mutations included seven missense mutations, a 1 bp deletion, and a 12 bp insertion. Loss of heterozygosity at the AXIN1 locus was present in four of five informative HCCs with AXIN1 mutations, suggesting a tumor suppressor function of this gene. Park et al. (2005) showed that mutations of AXIN 1 are late events in hepatocellular carcinogenesis.
Entity name
Medulloblastoma
Note
To find out if Axin is also involved in the pathogenesis of sporadic medulloblastomas, Dahmen et al. (2001) analyzed 86 cases and 11 medulloblastoma cell lines for mutations in the AXIN1 gene. Using single-strand conformation polymorphism analysis, screening for large deletions by reverse transcription-PCR, and sequencing analysis, a single somatic point mutation in exon 1 (Pro255Ser) and seven large deletions (12%) of AXIN1 were detected. Baeza et al. (2003) screened 39 sporadic cerebellar medulloblastomas for alterations in the AXIN1 gene. The authors found missense AXIN1 mutations in two tumours, CCC-->TCC at codon 255 (exon 1, Pro-->Ser) and TCT-->TGT at codon 263 (exon 1, Ser-->Cys). Furthermore, the A allele at the G/A polymorphism at nucleotide 16 in intron 4 was significantly over-represented in medulloblastomas (39 cases; G 0.76 vs-A 0.24) compared to healthy individuals (86 cases; G 0.91 vs A 0.09; P=0.0027). Yokota et al. (2002) showed another AXIN1 mutation in exon 3, corresponding to GSK-3beta binding site.
Entity name
Colorectal carcinoma
Note
Hart et al. (1998) report on overexpression of Axin1 in connection to the downregulation of wild-type beta-catenin in colon cancer cells. In addition, Axin1 dramatically facilitated the phosphorylation of APC and beta-catenin by GSK3 beta in vitro. Another group (Jin et al., 2003) analyzed 54 colorectal cancer tissues for mutations in AXIN1 gene. They found 3 silent mutations, 6 missense point mutations in different functionally important regions. The missense mutation rate was hence 11%, suggesting that Axin 1 deficiency may contribute to the onset of colorectal tumorigenesis. Segditsas and Tomlinson (2006) report on mutations in AXIN1 in microsatellite-unstable colon cancers. Three AXIN1 missense variants P312T, R398H, and L445M were detected in 1 of 124 patients with multiple colorectal adenomas. Three other missense mutations, D545E, G700S, and R891Q, were found. The overall frequency of the rare variants was significantly higher in the patients as compared with the controls (Fearnhead et al., 2004).
Entity name
Brain tumors
Note
A sample of 72 neuroepithelial brain tumors was investigated for AXIN-1 gene changes by Nikuseva Martic et al. (2010). Polymorphic marker for AXIN-1, showed loss of heterozygosity in 11.1% of tumors. Down regulation of axin expression and up regulation of beta-catenin were detected. Axin was observed in the cytoplasm in 68.8% of samples, in 28.1% in both the cytoplasm and nucleus and 3.1% had no expression. Comparison of mean values of relative increase of axin and beta-catenin showed that they were significantly reversely proportional (P=0.014) in a set of neuroepithelial brain tumors. Pecina-Slaus et al. (2011) also explored axins existence at the subcellular level in glioblastomas and showed that the highest relative quantity of axin was measured when the protein was in the nucleus and the lowest relative quantity of axin when the protein was localized in the cytoplasm.
Entity name
Ovarian endometroid adenocarcinomas
Note
Wu et al. (2001) report on a nonsense mutation in one ovarian endometroid adenocarcinoma (OEA). They also found another missense AXIN1 sequence alteration in OEA-derived cell lines.
Entity name
Lung cancer
Note
In 105 lung SCC and adenocarcinoma tissue samples, the cytoplasmic expression of Axin was significantly lower than in normal lung tissues. Western blot analysis also demonstrated that the relative expression quantity of Axin was significantly reduced in lung cancer tissues compared with normal lung tissues. Nuclear expression of Axin was observed in 21 cases (20%) of lung cancers (Xu et al., 2011).
Entity name
Oesophageal squamous cell carcinoma
Note
Nakajima et al. (2003) found reduced expression of Axin1 in oesophageal squamous cell carcinoma. Several mutations have also been reported in oesophageal squamous cell carcinoma.
Entity name
Cervical cancer
Note
Su et al. (2003) examined AXIN1 in cervical cancer. Among the 30 tested cervical cancers mutation analysis of AXIN1 revealed that one specimen had a heterozygous mutation at codon 740. Six polymorphisms were also found. Immunohistochemistry showed no relationship between the protein expression patterns and mutation of AXIN1.
Entity name
Prostate cancer
Note
Yardy et al. (2009) reported on AXIN1 mutations in advanced prostate cancer. They found 7 mutations in prostate cancer cases and 4 polymorphisms in prostate cancer cell lines.
Entity name
Caudal duplication anomaly
Note
Hypermethylation of the AXIN1 promoter is associated with the caudal duplication anomalies. Oates et al. (2006) examined methylation at the promoter region of the AXIN1 gene in monozygotic twins. The promoter region of the AXIN1 gene was significantly more methylated in the twin with the caudal duplication than in the unaffected twin.
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 12072559 | 2002 | Subcellular distribution of Wnt pathway proteins in normal and neoplastic colon. | Anderson CB et al |
| 12555076 | 2003 | AXIN1 mutations but not deletions in cerebellar medulloblastomas. | Baeza N et al |
| 19141611 | 2009 | Identification of zinc-finger BED domain-containing 3 (Zbed3) as a novel Axin-interacting protein that activates Wnt/beta-catenin signaling. | Chen T et al |
| 20846493 | 2010 | Coexpression and protein-protein complexing of DIX domains of human Dvl1 and Axin1 protein. | Choi SH et al |
| 12781135 | 2003 | A role of Dishevelled in relocating Axin to the plasma membrane during wingless signaling. | Cliffe A et al |
| 14981260 | 2004 | Nuclear-cytoplasmic shuttling of Axin regulates subcellular localization of beta-catenin. | Cong F et al |
| 11585731 | 2001 | Deletions of AXIN1, a component of the WNT/wingless pathway, in sporadic medulloblastomas. | Dahmen RP et al |
| 19909509 | 2009 | The structural and functional determinants of the Axin and Dishevelled DIX domains. | Ehebauer MT et al |
| 15520370 | 2004 | Multiple rare variants in different genes account for multifactorial inherited susceptibility to colorectal adenomas. | Fearnhead NS et al |
| 19390532 | 2009 | Axin localizes to the centrosome and is involved in microtubule nucleation. | Fumoto K et al |
| 11438668 | 2001 | Axin facilitates Smad3 activation in the transforming growth factor beta signaling pathway. | Furuhashi M et al |
| 9601641 | 1998 | Downregulation of beta-catenin by human Axin and its association with the APC tumor suppressor, beta-catenin and GSK3 beta. | Hart MJ et al |
| 14566817 | 2003 | Detection of point mutations of the Axin1 gene in colorectal cancers. | Jin LH et al |
| 18632848 | 2008 | SUMOylation target sites at the C terminus protect Axin from ubiquitination and confer protein stability. | Kim MJ et al |
| 20858899 | 2010 | The protein stability of Axin, a negative regulator of Wnt signaling, is regulated by Smad ubiquitination regulatory factor 2 (Smurf2). | Kim S et al |
| 19731416 | 2009 | Axin determines cell fate by controlling the p53 activation threshold after DNA damage. | Li Q et al |
| 15473860 | 2004 | The Wnt signaling pathway in development and disease. | Logan CY et al |
| 15067197 | 2004 | Axin: a master scaffold for multiple signaling pathways. | Luo W et al |
| 15579909 | 2005 | Axin contains three separable domains that confer intramolecular, homodimeric, and heterodimeric interactions involved in distinct functions. | Luo W et al |
| 12771989 | 2003 | Reduced expression of Axin correlates with tumour progression of oesophageal squamous cell carcinoma. | Nakajima M et al |
| 19633924 | 2010 | Changes of AXIN-1 and beta-catenin in neuroepithelial brain tumors. | Nikuseva Martić T et al |
| 21087614 | 2011 | Critical scaffolding regions of the tumor suppressor Axin1 are natively unfolded. | Noutsou M et al |
| 16773576 | 2006 | Increased DNA methylation at the AXIN1 gene in a monozygotic twin from a pair discordant for a caudal duplication anomaly. | Oates NA et al |
| 15698401 | 2005 | Mutations of beta-catenin and AXIN I genes are a late event in human hepatocellular carcinogenesis. | Park JY et al |
| 21648318 | 2011 | AXIN-1 protein expression and localization in glioblastoma. | Pećina-Slaus N et al |
| 10921899 | 2000 | Wnt signaling and cancer. | Polakis P et al |
| 12717450 | 2003 | A role for Wnt signalling in self-renewal of haematopoietic stem cells. | Reya T et al |
| 15735151 | 2005 | The links between axin and carcinogenesis. | Salahshor S et al |
| 10700176 | 2000 | AXIN1 mutations in hepatocellular carcinomas, and growth suppression in cancer cells by virus-mediated transfer of AXIN1. | Satoh S et al |
| 17143297 | 2006 | Colorectal cancer and genetic alterations in the Wnt pathway. | Segditsas S et al |
| 17554179 | 2007 | Crystallization and preliminary X-ray crystallographic studies of the axin DIX domain. | Shibata N et al |
| 12883680 | 2003 | Mutation analysis of CTNNB1 (beta-catenin) and AXIN1, the components of Wnt pathway, in cervical carcinomas. | Su TH et al |
| 12101426 | 2002 | Mutational spectrum of beta-catenin, AXIN1, and AXIN2 in hepatocellular carcinomas and hepatoblastomas. | Taniguchi K et al |
| 19038973 | 2009 | Identification of WNT/beta-CATENIN signaling pathway components in human cumulus cells. | Wang HX et al |
| 14630927 | 2004 | Nucleo-cytoplasmic shuttling of Axin, a negative regulator of the Wnt-beta-catenin Pathway. | Wiechens N et al |
| 9891778 | 1998 | Mechanisms of Wnt signaling in development. | Wodarz A et al |
| 11719457 | 2001 | Diverse mechanisms of beta-catenin deregulation in ovarian endometrioid adenocarcinomas. | Wu R et al |
| 21496867 | 2011 | Disabled-2 and Axin are concurrently colocalized and underexpressed in lung cancers. | Xu HT et al |
| 10196136 | 1999 | Phosphorylation of axin, a Wnt signal negative regulator, by glycogen synthase kinase-3beta regulates its stability. | Yamamoto H et al |
| 18514389 | 2009 | Mutations in the AXIN1 gene in advanced prostate cancer. | Yardy GW et al |
| 12209999 | 2002 | Role of Wnt pathway in medulloblastoma oncogenesis. | Yokota N et al |
| 9230313 | 1997 | The mouse Fused locus encodes Axin, an inhibitor of the Wnt signaling pathway that regulates embryonic axis formation. | Zeng L et al |
Other Information
Locus ID:
NCBI: 8312
MIM: 603816
HGNC: 903
Ensembl: ENSG00000103126
Variants:
dbSNP: 8312
ClinVar: 8312
TCGA: ENSG00000103126
COSMIC: AXIN1
RNA/Proteins
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 38183076 | 2024 | R-spondin-1 induces Axin degradation via the LRP6-CK1ε axis. | 0 |
| 38715121 | 2024 | UBE2N promotes cell viability and glycolysis by promoting Axin1 ubiquitination in prostate cancer cells. | 0 |
| 38972426 | 2024 | Hypoxia exposure induces lactylation of Axin1 protein to promote glycolysis of esophageal carcinoma cells. | 0 |
| 38183076 | 2024 | R-spondin-1 induces Axin degradation via the LRP6-CK1ε axis. | 0 |
| 38715121 | 2024 | UBE2N promotes cell viability and glycolysis by promoting Axin1 ubiquitination in prostate cancer cells. | 0 |
| 38972426 | 2024 | Hypoxia exposure induces lactylation of Axin1 protein to promote glycolysis of esophageal carcinoma cells. | 0 |
| 36510340 | 2023 | Association of AXIN1 rs12921862 C/A and rs1805105 G/A and CTSB rs12898 G/A polymorphisms with papillary thyroid carcinoma: A case-control study. | 1 |
| 37029301 | 2023 | SUMOylation of RNF146 results in Axin degradation and activation of Wnt/β-catenin signaling to promote the progression of hepatocellular carcinoma. | 5 |
| 37201672 | 2023 | RSK2 inactivation cooperates with AXIN1 inactivation or β-catenin activation to promote hepatocarcinogenesis. | 1 |
| 37380960 | 2023 | Association between AXIN1 gene polymorphism (rs9921222) of WNT signaling pathway and susceptibility to osteoporosis in Egyptian patients: a case-control study. | 0 |
| 37548359 | 2023 | The Axin scaffold protects the kinase GSK3β from cross-pathway inhibition. | 2 |
| 37582359 | 2023 | AXIN1 bi-allelic variants disrupting the C-terminal DIX domain cause craniometadiaphyseal osteosclerosis with hip dysplasia. | 2 |
| 36510340 | 2023 | Association of AXIN1 rs12921862 C/A and rs1805105 G/A and CTSB rs12898 G/A polymorphisms with papillary thyroid carcinoma: A case-control study. | 1 |
| 37029301 | 2023 | SUMOylation of RNF146 results in Axin degradation and activation of Wnt/β-catenin signaling to promote the progression of hepatocellular carcinoma. | 5 |
| 37201672 | 2023 | RSK2 inactivation cooperates with AXIN1 inactivation or β-catenin activation to promote hepatocarcinogenesis. | 1 |
Citation
Nives Pecina-Slaus ; Tamara Nikuseva Martic ; Tomislav Kokotovic
AXIN1 (axin 1)
Atlas Genet Cytogenet Oncol Haematol. 2011-05-01
Online version: http://atlasgeneticsoncology.org/gene/379/axin1-(axin-1)
