Clinics and Pathology
Note
The most common childhood malignancy is acute lymphoblastic leukemia (ALL), representing approximately one-third of all newly diagnosed pediatric malignancies. Two types of ALL exist, those involving B cells and those involving T-cells, with T-cell ALL comprising 10-25% of all cases.
Phenotype stem cell origin
T-cells mature in the thymus orchestrate cellular immunity, and are integral to the process of humoral immunity provided by B-cells. Early thymic progenitors, or lymphoblasts, enter the thymus, and in their journey through this organ differentiate by the process of somatic rearrangement of T-cell receptor (TCR) genes. The completely differentiated T-cell must be both functional and self-tolerant, a stringent requirement which results in an approximately 95% attrition rate of mature T-cells. The approximately 5% that actually leave the thymus circulate in the blood as well as through lymphatic and non-lymphatic organs [Karrman and Johansson, 2017]. T-ALL is, therefore, a neoplasm of these lymphoblasts committed to the T-cell lineage.
Etiology
Because of the relatively low incidence of T-ALL compared with B-ALL, there is more data specifically regarding the etiology of B-ALL; however, the two diseases may share causative mechanisms. Both environmental exposure and genetic predisposition have been cited as possible etiologies for ALL, with exposure to ionizing radiation considered a known risk factor. The inheritance of both rare high penetrance, and more common low penetrance alleles, have been suggested by several studies, with inherited polymorphisms of CDKN2A the only confirmed variant directly attributable to an increased risk of developing T-ALL. In addition, genetic predisposition has also been demonstrated in a few studies where families have shown a high concordance of T-ALL in siblings [Karrman and Johansson, 2017].
It is the progressive accumulation of both genetic and epigenetic changes involving the immature thymocyte within the thymus that leads to T-ALL. Approximately half of all patients with T-ALL present with chromosome rearrangements that involve T-cell receptor genes, most commonly T-cell receptors α and δ (TRA and TRD, respectively) at chromosome band 14q11, T-cell receptor β (TRB) at chromosome band 7q34, or T-cell receptor γ (TRG) at 7p14.1. Unlike non-malignant conditions that give rise to a polyclonal T-cell response, T-cell ALL arises from a single cell that gives rise to a clone with an identical T-cell receptor gene rearrangement. These rearrangements will juxtapose one of these T-cell receptor genes with many critical partner genes, many of which code for transcription factors. This results in dysregulation of transcription of the partner gene, which is the main driver in leukemogenesis. Some of the most common partner genes include HOX11 (TLX1) at 10q24, HOX11L2 (TLX3) at 5q35, MYC at 8q24.1, TCL1 at 14q32, TAL1 at 1p32, LMO1 at 11p15, LMO2 at 11p13, and LYL1 at 19p13. As many of these rearrangements are submicroscopic, they cannot routinely be detected by conventional cytogenetic analysis, but require fluorescence in situ hybridization (FISH) to confirm the diagnosis. Approximately 20% of cases will also demonstrate simultaneous rearrangement of IGH@ [Borowitz and Chan, 2008].
DNA sequence mutations and copy number variants are also identified in T-ALL and are believed to be significant contributors to leukemogenesis. These mutations are found in genes involved in the JAK-STAT and Ras/PI3K/AKT pathways, in epigenetic regulation, in mRNA maturation and ribosome activity, in modifying histone methylation or acetylation, and that function as regulators of transcription. In fact, mutations in epigenetic regulators are found in over 50% of pediatric T-ALL cases, making this a common mechanism of tumorigenesis. Activating mutations of NOTCH1 (occurring in at least 60% of T-ALL cases) and loss-of-function mutations in FBXW7 are also commonly found in T-ALL and result in inhibition of ubiquitin-mediated degradation of the activated form of NOTCH1 [Iacobucci and Mullighan, 2017; Karrman and Johansson, 2017].
Epidemiology
Approximately 6000 new cases of ALL are diagnosed each year in the United States. T-ALL can occur in children or adults, but is more common in the pediatric population and more often diagnosed in adolescents than in young children, with a median age of onset of 9 years, compared with the more common precursor B-cell ALL which has an incidence peak between 2 and 5 years. In addition, there is a marked male predominance, with boys having a threefold-increased risk of developing T-ALL compared with females. It is unclear why T-ALL is primarily a disease of older children and why males are more preferentially affected compared with B-ALL [Karrman and Johansson, 2017].
Clinics
Both T-ALL and T-cell lymphoblastic lymphoma (T-LBL) are aggressive and highly heterogenous diseases which originate from genetic and epigenetic alterations in immature thymocytes during the process of differentiation within the thymus. T-ALL and T-LBL, which demonstrate overlapping clinical, morphological and immunophenotypic features, have been considered different manifestions of the same disease, with the only difference being the site of presentation. T-ALL patients present with extensive bone marrow involvement and lymphoblasts in peripheral circulation, a mediastinal mass in two-thirds of patients, splenomegaly, adenopathy, and central nervous system involvement. T-LBL is primarily confined to a mediastinal mass lesion with minimal or no bone marrow involvement. [Burkhardt 2010; Karrman and Johansson, 2017]. However, subtle immunophenotypic, molecular and cytogenetic differences suggest that T-ALL and T-LBL may, in fact, be biologically distinct diseases [Burkhardt 2010; Basso et al, 2011].
T-ALL can present acutely or with symptoms that develop and progress over several months. Proliferation of the malignant clone in different tissues accounts for the common clinical features of T-ALL. Expansion of the clone within the bone marrow results in a high leukocyte count and a concurrent suppression of normal hematopoiesis, resulting in deficiency of normal peripheral blood cells, especially thrombocytes. Symptoms include fever, joint and bone pain, recurrent infection, lethergy, paleness, mucosal bleeding, and hepatosplenomegaly. CNS involvement results in neurological symptoms including headache, visual impairment, and nausea. The mediastinal mass seen in approximately 50-60% of T-ALL patients can cause superior vena cava syndrome. The mediastinal mass(s) found in T-LBL predomininately involve the anterior mediastinum and cause progressive dyspnea with cough, edema, and elevated jugular venous pressure. While the clinical manifestations are somewhat different, the only true distinguishing feature between T-ALL and T-LBL is the blast count in bone marrow (>25% blasts in T-ALL). [Burkhardt, 2010]
Pathology
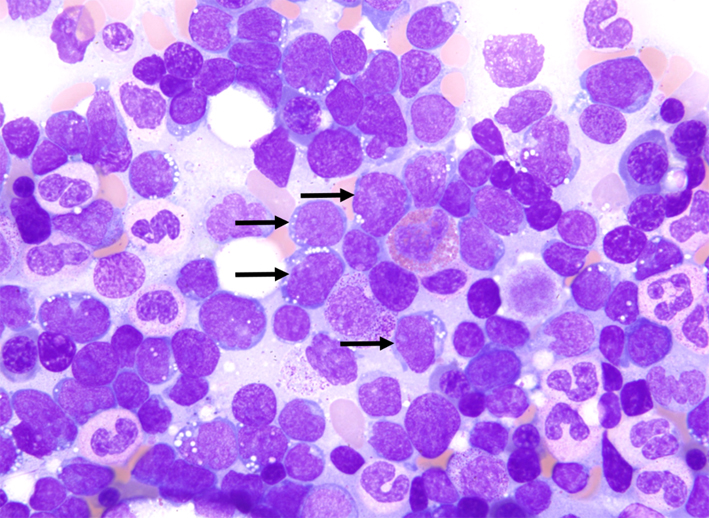
The lymphoblasts in B-ALL and T-ALL are morphologically indistinguishable from one another, necessitating the use of immunophenotyping for lineage determination. ALL blasts are generally of intermediate size (up to twice the size of a small lymphocyte) with a high nuclear/cytoplasmic ratio, although there can be variation in size range. The blasts demonstrate scant, basophilic cytoplasm with homogenous and somewhat condensed chromatin. Nucleoli may or may not be present depending on the size of the blast, as might cytoplasmic vacuoles. When the blasts resemble more mature lymphocytes, immunophenotyping is required to distinguish T-ALL from a mature (peripheral) T-cell leukemia [Borowitz and Chan, 2008; Onciu M, 2007].
As mentioned, immunophenotyping by immunohistochemistry and/or flow cytometry is critical in cases of ALL to determine the cell lineage (B-cell vs. T-cell). All T-cell ALLs express CD45 and cytoplasmic CD3 (CD3 may also be weakly expressed on the cell surface), with CD3 being T-cell lineage specific and thus an important maker for lineage differentiation. Other T-cell antigens consistently expressed include CD2, CD5, and CD7. The DNA polymerase terminal deoxynucleotidyl transferase (TdT) is expressed in 80-90% of T-ALL cases. In addition, variable expression of CD1, CD4, CD8, CD10, and CD56 is observed. Lineage-inappropriate expression of myeloid markers including CD11b, CD13, CD33, and CD66c are also observed; however, their expression is of no prognostic significance and do not necessarily warrant a diagnosis of biphenotypic leukemia [Onciu M, 2007; Karrman and Johansson, 2017].
Because the expression of CDs change with the maturation stage of thymocytes, the array of CD markers in a case can indicate the stage at which the differentiation block occurred. The stages of differentiation include: 1) pro-T (cCD3+, CD7+, CD2-, CD1a-, CD34+/-); 2) pre-T (cCD3+, CD7+, CD2+, CD1a-, CD34+/-); 3) cortical T (cCD3+, CD7+, CD2+, CD1a+, CD34-); and 4) medullary T (cCD3+, CD7+, CD2+, CD1a-, CD34-, surface CD3+) [Borowitz and Chan, 2008]. Cases of pro- and pre T-ALL tend to have a more inferior outcome compared with those designated as cortical or medullary T-ALL. The recently delineated category of T-ALL known as early T-cell precursor ALL (ETP-ALL) has a distinct immunophenotype. This T-ALL demonstrates an immature immunophenotype, expresses myeloid and/or stem cell markers, and has a poor outcome; however, recent studies suggest intensive treatment can overcome the poor prognostic outcome in some patients [Belver and Ferrando, 2016; Karrman and Johansson, 2017].
This bone marrow aspirate from a 15-year-old boy with 70% marrow involvement by blasts. The blasts (black arrows) were medium size with a high nuclear/cytoplasmic ratio and demonstrated fine chromatin, occasional nucleoli and cytoplasmic vacuoles. Giemsa, 1000x
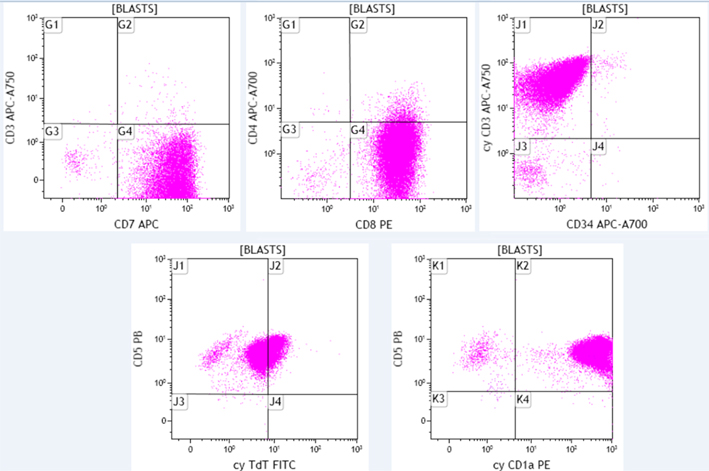
The blast population from the same patient expressed the following antigens: CD4 (variable), CD5, CD7, CD8, cytoplasmic CD3, cytoplasmic TdT (dim) and cytoplasmic CD1a. They lacked expression of surface CD3 and CD34. Additionally, the cells expressed CD2, CD10, CD38, and CD45 (dim) which are not shown here. The immunophenotype is consistent with T-acute lymphoblastic leukemia.
Treatment
Patients with T-cell ALL generally present with more high-risk features than do those with B-ALL. These features include an increased resistance to chemotherapy, a tendency for earlier relapse, and CNS involvement. Therefore, most patients are treated aggressively, including intrathecal chemotherapy and CNS radiation therapy. The outcome of children with T-ALL has improved with intensification of multimodal therapy. Chemotherapy regimens in children with T-ALL include initial induction therapy for 4-6 weeks followed by intensive combination therapy for 6-8 months and low-intensity anti-metabolite-based therapy for 18-30 months. Induction therapy includes vincristine, a glucocorticoid such as dexamethasone, L-asparaginase, and possibly an anthracycline such as daunorubicin [Hunger and Mullighan, 2015].
Allogeneic hematopoietic stem cell transplantation has been found to be an effective treatment for high-risk T-ALL patients; however, relapse is not uncommon in such cases, reducing its curative potential. Continuous monitoring for minimal residual disease following transplantation, using various molecular methods including qPCR to evaluate for fusion transcripts such as SIL/TAL1, is critical so that immediate intervention can be provided at an early stage of relapse [Zhao et al, 2017].
Prognosis
The improvement in outcome of children with T-ALL continues to lag behind its B-cell counterpart. With risk-stratified consolidated chemotherapy, 5-year event free survival for B-ALL stands at 80% and overall survival at 90%. However, 5-year event free survival and overall survival for T-ALL is just over 70% and 80%, respectively. Patients with T-ALL have more induction failures and extramedullary relapses than those with B-ALL. Relapsed T-ALL occurs in up to 25% of children and is associated with a much poorer prognosis (30-50% survival). Therefore, monitoring for minimal residual disease (MRD), usually by qPCR analysis of TCR genes is critical to evaluate the effectiveness of treatment [Pui CH et al, 2015; Karrman and Johansson, 2017]. If a child continues to have a high level of MRD at the end of induction, the risk of relapse is high [Chu et al, 2017]. Recent genomic studies have begun to uncover the basis for relapse in this disease, which appears to involve both clonal evolution and selection of genetic variants that drive resistence. One such variant involves mutation in the cytosolic 5-nucleotidase II (
NT5C2) gene which are found in 20% of patients with relapsed T-ALL [Tzoneva et al, 2013]. In addition, the
TFDP3 gene has been found to confer chemoresistance in children with T-ALL with MRD [Chu et al, 2017].
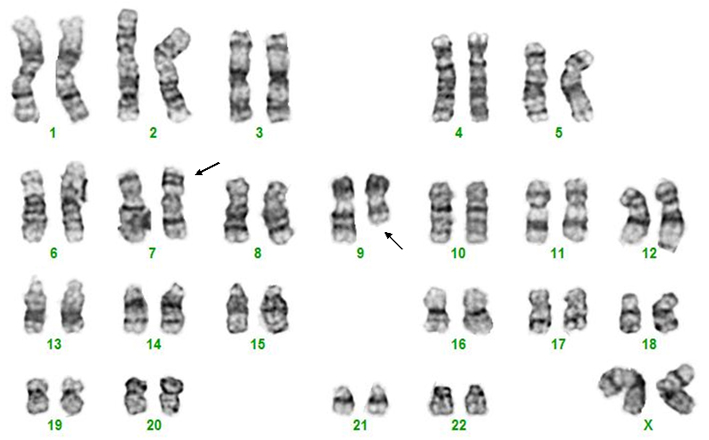
Bone marrow cytogenetic evaluation reveals a pericentric inversion of chromosome 7 as well as a del(9q) chromosome in a child with T-ALL.
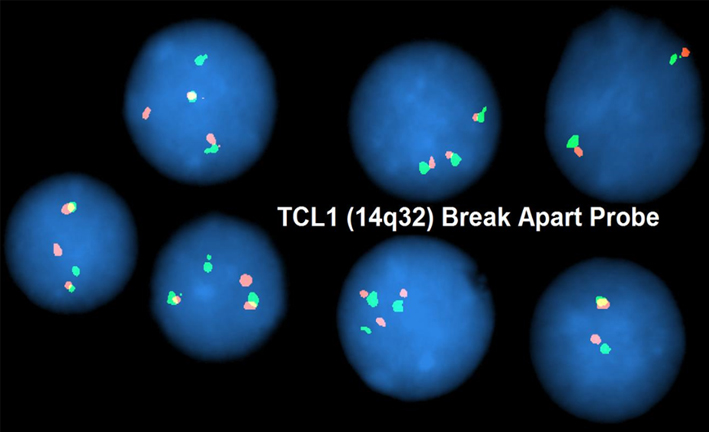
FISH analysis utilizing a TCL1 breakapart probe reveals gene rearrangement identified by seperate red and green signals in a patient with T-ALL. The karyotype also included an additional copy of chromosome 14.
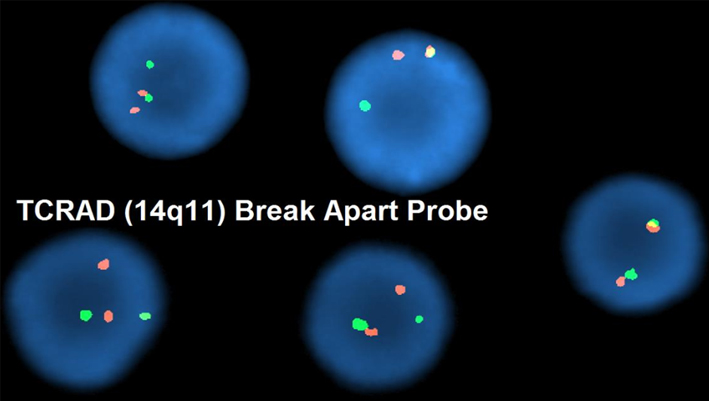
FISH analysis utilizing a TCR alpha/delta breakapart probe reveals rearrangement of the locus in a patient with T-ALL.
Genes Involved and Proteins
Note
While many of the cytogenetic and molecular abnormalities characteristic of T-cell ALL have been known for some time, gene expression profiling and various sequencing strategies including exome and whole genome sequencing, as well as transcriptome analysis, have recently increased our knowledge of the molecular pathogenesis of this disease. Over 100 genes are now known to be mutated in T-ALL, with only NOTCH1 and CDKN2A mutations considered to be common events. Recent sequencing studies have demonstrated that each case of T-ALL carries 10-20 biologically relevant mutations, suggesting one or a few primary mutations and a number of secondary changes that impair the normal differentiation and proliferation of T-cells. The pattern of these genomic changes is not random, and certain specific combinations of genomic mutations are found, suggesting an interrelationship of the genes responsible for T-cell leukemogenesis. In addition, recent studies have uncovered mutations in non-coding regions of DNA that alter transcription factors or result in misexpression of micro RNAs and long non-coding RNAs, underscoring the fact that there is much still to be learned about the pathogenesis of T-ALL [Girardi et al, 2017].
Excellent recent reviews of the genes and molecular mechanisms involved in T-cell acute lymphoblastic leukemia are provided in Belver and Ferrando, 2017 and Liu et al, 2017. Much of the information provided below was abstracted from these two references, as well as the NCBI Reference Sequence Database (RefSeq). Additional references are provided as necessary.
Gene group
T-cell receptor genes
Gene name
TRA (T-cell receptor alpha locus)
Gene name
TRD (T-cell receptor delta locus)
Gene name
TRG (T-cell receptor gamma locus)
Gene group
Genes involved in the NOTCH1 Pathway
Gene name
FBXW7 (F-box and WD repeat domaining 7)
Protein description
This gene encodes a member of the F-box protein family, which constitute one of the four subunits of ubiquitin protein ligase complex that functions in phosphorylation-dependent ubiquitination. This protein binds directly to cyclin E and probably targets cyclin E for ubiquitin-mediated degradation. [RefSeq, Mar 2012]. Inactivating mutations of FBXW7 are found in 8-30% of T-ALL cases and appear to increase the levels of NOTCH1 and MYC proteins.This gene encodes a member of the NOTCH family of proteins. Members of this Type I transmembrane glycoprotein family share structural characteristics including an extracellular domain consisting of multiple epidermal growth factor-like (EGF) repeats [RefSeq, Jan 2016]. Notch signaling is an evolutionarily conserved intercellular signaling pathway that transduces extracellular signals to affect nuclear genes leading to transcriptional changes. NOTCH1 acts as a transcriptional activator, regulating the expression of genes involved in proliferation, metabolism, and cell cycling such as MYC and
CCND3. Constitutive activation of NOTCH1 occurs less commonly through the t(7;9)(q34;q34.3) and more commonly through activating mutations, found in 50% of T-ALL cases. Some studies have demonstrated favorable early response in children with NOTCH1 mutations; however, not all studies have shown improved overall survival [Karrman and Johansson, 2017].
Gene group
Oncogenes that function as transcription factors
Gene name
HOXA@ (Homeobox A cluster 9/10/11))
Protein description
The Hox family of homeodomain transcription factors plays an important role in regulating hematopoiesis. Activation of HOXA genes occurs often through juxtaposition with the TRB gene in either the t(7;7)(p15.3;q34) or inv(7)(p15.3q34) (occurring in 3% of T-ALL cases), but other molecular mechanisms leading to HOXA dysregulation have recently been identified [Bond et al, 2016]. The prognostic impact is unknown.
Protein description
This locus encodes a transcriptional regulator that contains two cysteine-rich LIM domains but lacks a DNA-binding domain. LIM domains may play a role in protein interactions. Thus the encoded protein may regulate transcription by competitively binding to specific DNA-binding transcription factors [RefSeq, Jul 2012]. Chromosome rearrangements between this locus and either the TRB or TRD gene have been observed in 1% T-ALL cases.
Protein description
LMO2 encodes a cysteine-rich, two LIM-domain protein that is required for yolk sac erythropoiesis. The LMO2 protein has a critical role in hematopoietic development and is highly conserved. The transcription start site is located approximately 25 kb downstream from the 11p13 T-cell translocation cluster where a number of T-cell ALL specific translocations occur [RefSeq, Nov 2008]. Depending on the specific cytogenetic abnormality, the LMO2 gene is involved in 1-6% of T-ALL cases.
Gene name
LYL1 (LYL1, basic helix-loop-helix family member)
Protein description
This gene codes for a basic helix-loop-helix transcription factor that plays a role in blood vessel maturation and in hematopoeisis [RefSeq, Sep 2010]. Disruption is caused by the t(7;19)(q34;p13.2) which involves the TRB gene, and is observed in 1% of T-cell ALL cases.
Gene name
MLLT10 (MLLT10, histone lysine methyltransferase DOT1L cofactor)
Protein description
This gene encodes a transcription factor. It fuses with the PICALM gene at 11q14 resulting in a specific gene expression profile involving upregulation of HOXA genes (HOXA5 and HOXA9) that is associated with a poorer prognosis.
Gene name
MYB (MYB proto-oncogene, transcription factor)
Protein description
The MYB protein functions as a transcription regulator and plays an essential role in the regulation of hematopoiesis. Upregulation of MYB through a chromosome 6q deletion or the t(6;7)(q23 q34) which occurs in 3% of all T-ALL cases and in 10% of very young T-ALL patients
Gene name
MYC (MYC proto-oncogene, bHLH transcription factor)
Protein description
This gene is a proto-oncogene that encodes a nuclear phosphoprotein that plays a role in cell cycle progression, apoptosis and cellular transformation in many cancers. The encoded protein regulates the transcription of specific target genes [RefSeq, Aug 2017]. MYC is also an important mediator of NOTCH1 induced transformation is dysregulated in the t(8;14)(q24.1;q11.2) which involves the TRA gene, and generally predicts a poor prognosis in T-ALL.
Protein description
The NKX2-1 homeobox gene is one of a number of genes including TLX1, TLX3, and
NKX2-5 which encode transcription factors that are physiologically silent during hematopoiesis, but which undergo ectopic activation in transformed thymocytes [Nagel et al, 2017]. T-ALLs with NKX2-1 mutations share a gene expression signature similar to cases with TLX1 expression and show arrest of an early cortical stage of thymocyte development. Aberrant expression of the gene occurs with the inv(14)(q11.2q13), inv(14)(q13q32.33), and t(7;14)(q34;q13), and is found in 5% of pediatric T-ALL cases.
Gene name
TAL1 (TAL bHLH transcription factor 1, erythroid differentiation factor)
Protein description
The TAL1 protein is a basic helix-loop-helix transcription factor that plays an essential role in regulating normal hematopoiesis. Upregulation of the TAL1 gene occurs by means of submicroscopic chromosome 1p deletion, insertions, or disruption of chromatin structure resulting in SIL-TAL1 fusion. Recent evidence demonstrates that TAL1 is, in fact, a master transcription factor in the core regulatory circuit which includes TAL1, LMO2, RUNX1,
GATA2, and
FLI1 genes that function to reinforce and stabilize gene expression programs via interlocking regulatory loops [Sanda and Leong, 2017]. Translocations of TAL1 are found in 1-3% of T-ALL cases while deletions are identified in up to 25% of cases. The prognostic significance of the SIL-TAL1 fusion gene is unclear, with some studies demonstrating a poorer outcome while others showed no significance (Karrman and Johannson, 2017].
Gene name
TAL2 (TAL bHLH transcription factor 2)
Protein description
This intronless gene encodes a helix-loop-helix protein and is activated by the t(7;9)(q34;q32) involving the TRB gene, found in about 1% of pediatric T-ALL cases.
Gene name
TLX1 (HOX11) (T-cell leukemia homeobox 1)
Protein description
This gene encodes a nuclear transcription factor that belongs to the NK-linked or NK-like (NKL) subfamily of homeobox genes. The TLX1 protein is required for normal development of the spleen during embryogenesis and specifies neuronal cell fates [RefSeq, Sep 2010]. The t(10;14)(q24;q11.2) juxtaposes the TLX1 gene with the promoter elements of TRA/TRD and is found in 5-10% of pediatric T-ALL cases. There appears to be no prognostic impact in pediatric T-ALL.
Gene name
TLX3 (HOX11L2) (T-cell leukemia homeobox 3)
Protein description
The TLX3 protein is an orphan homeobox protein that encodes a DNA-binding nuclear transcription factor. Upregulation of TLX3 through juxtaposition of the gene with strong T-cell regulatory elements in the BCL11B locus at 14q32 occurs in 20-25% of pediatric T-ALL cases and is associated with an inferior outcome.
Gene group
Tumor suppressor genes that function as transcription factors
Gene name
BCL11B (B-cell CLL/lymphoma 11b)
Protein description
This gene encodes a C2H2-type zinc finger protein. Although the specific function of this gene has not been determined, the protein is known to be a transcriptional repressor [RefSeq, Aug 2013]. Inactivating mutation or deletion of BCL11B is found in 10% of T-ALL cases. There appears to be no prognostic impact in pediatric T-ALL.
Protein description
This gene encodes an ETS family transcription factor required for development of hematopoietic stem cells Inactivating mutation or deletion of ETV6 is found in 13% of T-ALL cases.
Gene name
GATA3 (GATA binding protein 3)
Protein description
This gene encodes a protein which belongs to the GATA family of transcription factors. The protein is an important regulator of T-cell differentiation and plays a crucial role in the development of early T-cell progenitors. The gene is mutated in early T-cell precursor T-ALL. Inactivating mutation or deletion is observed in 5% of T-ALL cases.
Gene name
LEF1 (Lymphoid enhancer binding factor 1)
Protein description
This gene encodes a transcription factor that binds to a functionally important site in the T-cell receptor-alpha enhancer, thereby conferring maximal enhancer activity, and is a key mediator of WNT signaling. T-ALL with LEF1 mutations demonstrate high levels of MYC expression and appear to arrest at the early cortical thymocyte stage. Inactivating mutation or deletion of LEF1 is found in 10-15% of T-ALL cases.
Gene name
RUNX1 (Runt related transcription factor 1)
Protein description
The protein encoded by this gene represents the alpha subunit of core binding factor (CBF), a heterodimeric transcription factor that binds to the core element of many enhancers and promoters. CBF and is thought to play a prominent role in the development of normal hematopoiesis [RefSeq, Jul 2008]. Inactivating mutation or deletion of RUNX1 is found in 10-20% of T-ALL cases.
Protein description
This gene encodes a transcription factor that appears to play a role in hematopoiesis to maintain primitive stem cell quiescence and to promote differentiation of mature progenitors. Inactivating mutation or deletion of WT1 is found in 10% of T-ALL cases.
Gene group
Cell cycle regulatory genes
Gene name
CDKN2A (Cyclin dependent kinase inhibitor 2A)
Protein description
Inactivation of CDKN2A is believed to be a primary abnormality found in 88% of younger-age children with T-ALL and to be directly involved in T-cell leukemogenesis. This gene encodes protein isoforms including, p16
INK4A and p14
ARF, that function as negative regulators of the cell cycle by inhibiting CDK4 kinase to inhibit cell cycle G1 progression. Another specific isoform functions to stabilize
TP53 as it can interact with, and sequester, the E3 ubiquitin-protein ligase
MDM2, a protein responsible for the degradation of p53 [RefSeq, Sep 2012]. Chromosome 9p deletions can inactivate CDKN2A, an abnormality found in about 30% of pediatric T-ALL cases by conventional analysis and in 80% of cases by FISH. In approximately 50% of these patients, the deletion involves both chromosome 9 homologues, often through segmental uniparental disomy. CDKN2A loss often occurs together with activating mutations of NOTCH1.
Gene name
CDKN2B (Cyclin dependent kinase inhibitor 2B)
Protein description
This gene lies adjacent to CDKN2A in a region that is frequently mutated and deleted in a wide variety of tumors. The gene encodes a cyclin-dependent kinase inhibitor that prevents the activation of CDK kinases, resulting in negative regulation of cell cycle G1 progression [RefSeq, Jul 2008]. Chromosome 9p deletions resulting in loss of CDKN2B as well as CDKN2A are observed in about 30% of pediatric T-ALL cases by conventional analysis and in 80% of cases by FISH. In approximately 50% of these patients, the deletion involves both chromosome 9 homologues. Loss of CDKN2B is thought to be a secondary change.
Gene name
RB1 (RB transcriptional corepressor 1)
Protein description
The RB1 protein is a negative regulator of the cell cycle that stabilizes constitutive heterochromatin to maintain overall chromatin structure. The active, hypophosphorylated form of the protein binds transcription factor E2F to downregulate transcription of S phase genes [RefSeq, Jul 2008]. RB1 deletion is identified in 12% of pediatric T-ALL cases.
Gene group
Assembly protein
Gene name
PICALM (Phosphatidylinositol binding clathrin assembly protein)
Protein description
This gene encodes a clathrin assembly protein which recruits clathrin and adaptor protein complex 2 (AP2) to cell membranes and may determine the amount of membrane to be recycled [RefSeq, May 2011]. The t(10;11)(p12;q14) fuses the MLLT10 gene at 10p12 with the PICALM gene, resulting in a specific gene expression profile involving upregulation of HOXA genes (HOXA5, HOXA9, and HOXA19) found in 10% of pediatric T-ALL cases. The rearrangement is associated with a poorer prognosis.
Gene group
Signal transduction genes
Gene name
ABL1 (ABL proto-oncogene 1, non-receptor tyrosine kinase)
Protein description
This gene is a protooncogene that encodes a protein tyrosine kinase involved in cell division, adhesion, differentiation, and response to stress. The proteins SH3 domain negatively regulates its activity, whereby deletion of the region encoding this domain results in an oncogene. The ubiquitously expressed protein has DNA-binding activity regulated by CDC2-mediated phosphorylation, suggesting a cell cycle function [RefSeq, Aug 2014]. Amplification of 9q34 resulting from cryptic episomal NUP214-ABL1 translocation occurs in 6% of mostly pediatric T-ALL cases. This is an important diagnosis to make with either ABL1 or BCR/ABL1 FISH probe, as malignant clones with ABL1 amplification are sensitive to Gleevec®.
Gene name
FLT3 (fms related tyrosine kinase 3)
Protein description
This gene encodes a receptor tyrosine kinase that regulates hematopoiesis. This receptor protein is activated by binding of the fms-related tyrosine kinase 3 ligand to the extracellular domain resulting in autophosphorylation of the receptor. The activated receptor kinase then phosphorylates and activates multiple cytoplasmic effector molecules in pathways involved in apoptosis, proliferation, and differentiation of hematopoietic cells in bone marrow. [RefSeq, Jan 2015]. Activating mutations are identified in 5-10% of T-ALL cases.
Gene name
IL7R (Interleukin 7 receptor)
Protein description
The IL7R gene encodes a protein receptor for interleukin 7 (IL7) that has been shown to activate the JAK-STAT signalling pathway. The protein has been shown to play a critical role in V(D)J recombination during lymphocyte development. Inactivating mutations are found in 10% of T-ALL cases. Mutations of IL7R are particularily prevelant in ETP T-ALL.
Gene name
JAK1 (Janus kinase 1)
Protein description
This gene encodes a membrane protein tyrosine kinase. The kinase phosphorylates STAT proteins (signal transducers and activators of transcription) and plays a key role in interferon-alpha/beta and interferon-gamma signal transduction. [RefSeq, Mar 2016]. Activating mutations are found in 4-18% of T-ALL cases.
Gene name
JAK3 (Janus kinase 3)
Protein description
The gene encodes a tyrosine kinases involved in cytokine receptor-mediated intracellular signal transduction. It is predominantly expressed in immune cells and transduces a signal in response to its activation via tyrosine phosphorylation by interleukin receptors. [RefSeq, Jul 2008]. Activating mutations are found in 7% of T-ALL cases.
Gene name
NUP214 (Nucleoporin 214)
Protein description
The nuclear pore complex extends across the nuclear envelope and forms a gateway that regulates the flow of macromolecules between the nucleus and the cytoplasm. This gene is a member of the FG-repeat-containing nucleoporins, which are the main components of the nuclear pore complex in eukaryotic cells. The NUP214 protein is localized to the cytoplasmic face of the nuclear pore complex where it is required for proper cell cycle progression and nucleocytoplasmic transport [RefSeq, Dec 2015]. NUP214 fuses with ABL1 resulting in episomal ABL1 amplification observed in 5% of T-ALL cases. A SET-NUP214 fusion is also identified in T-ALL as the result of a 3Mb deletion between the SET locus at 9q34.11 and the NUP214 locus. This is found in 3% of T-ALL cases.
Gene name
PTEN (Phosphatase and tensin homolog)
Protein description
The PTEN protein is a phosphatidylinositol-3,4,5-trisphosphate 3-phosphatase that functions as a tumor suppressor by negatively regulating AKT/PKB signaling pathway . Inactivating mutations or deletions of PTEN occur in 10-15% of T-ALL cases.
Gene group
Genes involved in epigenetic regulation
Gene name
EED (Embryonic ectoderm development)
Protein description
This gene encodes a member of the Polycomb-group (PcG) family. PcG family members form multimeric protein complexes involved in maintaining the transcriptional repressive state of genes by histone deacetylation [RefSeq, Jul 2008]. Inactivating mutations and deletions are identified in 10% of T-ALL cases.
Gene name
EZH2 (Enhancer of zeste 2 polycomb repressive complex 2 subunit)
Protein description
This gene encodes a member of the Polycomb-group (PcG) family. PcG family members form multimeric protein complexes which are involved in maintaining the transcriptional repressive state of genes over successive cell generations. The protein associates with the embryonic ectoderm development protein, the
VAV1 oncoprotein, and the X-linked nuclear protein. [RefSeq, Feb 2011]. Inactivating mutations and deletions are identified in 10-15% of T-ALL cases.
Gene name
KDM6A (Lysine demethylase 6A)
Protein description
This gene encodes a protein which contains a JmjC-domain and catalyzes the demethylation of tri/dimethylated histone H3 [RefSeq, Apr 2014]. Inactivating mutations and deletions are identified in 5-15% of T-ALL cases.
Gene name
PHF6 (PHD finger protein 6)
Protein description
This gene is a member of the plant homeodomain (PHD)-like finger (PHF) family. It encodes a protein with two PHD-type zinc finger domains, indicating a potential role in transcriptional regulation that localizes to the nucleolus [RefSeq, Jun 2010]. Inactivating mutations and deletions are identified in 16% of pediatric T-ALL cases.
Gene name
SUZ12 (SUZ12 polycomb repressive complex 2 subunit)
Protein description
This zinc finger gene encodes a protein with a zinc finger domain in the C terminus of the coding region. Inactivating mutations and deletions are identified in 10% of T-ALL cases.
Gene group
Transcriptional coactivator gene
Gene name
KMT2A (Lysine methyltransferase 2a)/MLL (mixed lymphoid leukemia)
Protein description
This gene encodes a transcriptional coactivator that plays an essential role in regulating gene expression during hematopoiesis and hematopoiesis. The KMT2A protein contains multiple conserved functional domains. One of these domains, the SET domain mediates chromatin modifications associated with epigenetic transcriptional activation. Processing of the protein by the enzyme Taspase 1 splits it into two fragments. These fragments reassociate and further assemble into different multiprotein complexes that regulate the transcription of specific target genes, including many of the HOX genes [RefSeq, Oct 2010]. A t(11;19)(q23;p13) fuses KMT2A with
MLLT1 in 5% of T-ALL cases.
Gene group
Ribosomal protein gene
Gene name
RPL10 (Ribosomal protein L10)
Protein description
This gene encodes a ribosomal protein that is a component of the 60S ribosome subunit. Mutations are found in 6% of pediatric T-ALL cases