Identity

Abstract
PARP1 (poly(ADP-ribose) polymerase 1) is a nuclear protein involved in the regulation of various biological processes including apoptosis, DNA repair for the maintenance of genome integrity, epigenetic marking of chromatin, assembly of higher-order chromatin structures, transcriptional activation, differentiation, proliferation, and cell cycle. Particularly, due to its decisive role in several DNA repair pathways, the inhibition of PARP1 has emerged as a prominent therapeutic option in cancer treatment, by improving the efficiency of chemotherapeutics or radiation therapy.
DNA/RNA
Note
PARylation is a widely used process in eukaryotes. In eukaryotic species, the distribution of PARP proteins strictly follows the distribution of PARGs and at least one of the macrodomain proteins is also always present (Perina et al., 2014). On the other hand, PARP proteins are less common in bacteria and are thought to be acquired through horizontal gene transfer (Alemasova & Lavrik, 2019; Perina et al., 2014). In thermophilic archaeon Sulfolobus solfataricus, a protein with oligo(ADP-ribosyl) transferase activity was identified (Faraone-Mennella, Gambacorta, Nicolaus, & Farina, 1998) and in a number of dsDNA viruses have also been found to possess PARP homologues (Perina et al., 2014).
Based on the sequence homology, humans are assumed to express 17 defined PARPs (Vyas, Chesarone-Cataldo, Todorova, Huang, & Chang, 2013). PARP1, the first PARP purified and cloned from human, is a constitutive and the best studied member of the PARP family of proteins (Citarelli et al., 2010). The PARP1 gene is conserved in chimpanzee, Rhesus monkey, dog, cow, mouse, rat, chicken, zebrafish, fruit fly, mosquito, C.elegans, A.thaliana, rice, and frog (Table 1).
Table 1. Pairwise alignment of PARP1 gene (in distance from human) (HomoloGene:1222, NCBI).
| Gene Species | Gene Symbol | Identity (%) DNA |
| vs. P.troglodytes | PARP1 | 99,2 |
| vs. M.mulatta | PARP1 | 97,7 |
| vs. C.lupus | PARP1 | 88,4 |
| vs. B.taurus | PARP1 | 88,4 |
| vs. M.musculus | Parp1 | 86,5 |
| vs. R.norvegicus | Parp1 | 86,1 |
| vs. G.gallus | PARP1 | 75,2 |
| vs. X.tropicalis | parp1 | 72 |
| vs. D.rerio | parp1 | 69,5 |
| vs. D.melanogaster | Parp | 49,5 |
| vs. A.gambiae | AgaP_AGAP003230 | 53,1 |
| vs. C.elegans | pme-1 | 48,2 |
| vs. A.thaliana | PARP1 | 50,2 |
| vs. O.sativa | Os07g0413700 | 51,7 |
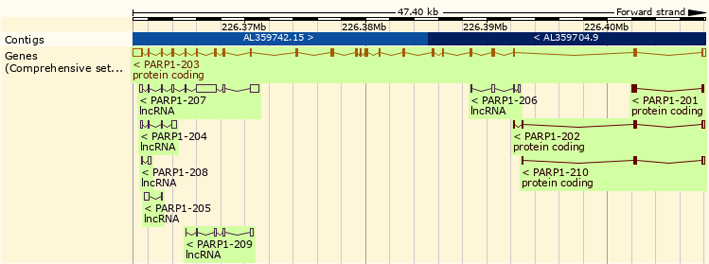
Description
Transcription
The human PARP1 promoter region does not contain typical regulatory elements, such as TATA or CAAT boxes. A near 40-base-pair region surrounding the transcription start site described as containing a near-consensus initiator element capable of initiating RNA polymerase II transcription (Abbotts & Wilson, 2017). Detailed analyses of the promoter regions of PARP1 genes in humans, rats, and mice showed that PARP1 promoter sequences have binding sites for transcription factors SP1, AP-2, YY1, ETS1, and NF1. In the distal promoter region of human PARP1 gene Candidate binding sites for several other factors including CDE, KLF4 (GKLF), BARB, RRM1 (MAZF), RREB1, HOX, GSX1 (GSH-1), CEBPB, NFIL3 (E4BP4), STAT6, cETSZ-1, PBX1, LEF1 (TCF), NF-kB, REL, ZNF148 (ZBP-89), KLF6 (CPBP), USF, CDF-1, EGR1, and IKZF1 (Ikaros 1) were also identified (Doetsch, Gluch, Poznanovic, Bode, & Vidakovic, 2012). Additionally, at the post-transcriptional level, miR-124, MIR223, let-7a, miR-7-5p, and MIR125B2 were shown to regulate cellular PARP1 expression (Dash et al., 2017; J. Lai et al., 2019; Wielgos et al., 2017).
Table 2. Transcripts of human PARP1 gene (Ensembl release 98 - September 2019)
| PARP1-201 | ENST00000366790.3 | 570 | - | - | Protein coding |
| PARP1-202 | ENST00000366792.3 | 553 | - | - | Protein coding |
| PARP1-203 | ENST00000366794.10 | 3978 | CCDS1554 | NM_001618.4 | Protein coding |
| PARP1-204 | ENST00000463968.5 | 830 | - | - | lncRNA |
| PARP1-205 | ENST00000468608.1 | 438 | - | - | lncRNA |
| PARP1-206 | ENST00000469663.1 | 542 | - | - | lncRNA |
| PARP1-207 | ENST00000490921.5 | 3165 | - | - | lncRNA |
| PARP1-208 | ENST00000491816.1 | 416 | - | - | lncRNA |
| PARP1-209 | ENST00000498787.1 | 628 | - | - | lncRNA |
| PARP1-210 | ENST00000629232.1 | 477 | - | - | Protein coding |
Pseudogene
Table 3. Pseudogenes of human PARP1 gene (GRCh38 Assembly, NCBI)
| Name/Gene ID | Description | Location (bp) | Aliases |
| PARP1P2 (ID: 145) | poly(ADP-ribose) polymerase 1 pseudogene 2 | Chr 14, NC_000014.9 (63123001..63123935) | ADPRTP2, PPOLP2 |
| PARP1P1 (ID: 144) | poly(ADP-ribose) +B7:D7polymerase 1 pseudogene 1 | Chr 13, NC_000013.11 (110936624..110940232) | ADPRTP1, PPOLP1 |
Proteins
Note
Table 4. Protein products of human PARP1 gene (Ensembl release 98 - September 2019)
| Name | Transcript ID | Protein | Charge | Isoelectric Point | Molecular Weight | CCDS | UniProt | RefSeq |
| PARP1-201 | ENST00000366790.3 | 155aa | 10,0 | 9,4862 | 17,324.99 g/mol | - | Q5VX85 | - |
| PARP1-202 | ENST00000366792.3 | 108aa | 3,0 | 7,7272 | 12,234.02 g/mol | - | Q5VX84 | - |
| PARP1-203 | ENST00000366794.10 | 1014aa | 31,5 | 9,3322 | 113,083.79 g/mol | CCDS1554 | A0A024R3T8 P09874 | NM_001618.4 |
| PARP1-210 | ENST00000629232.1 | 108aa | 3,0 | 7,7272 | 12,234.02 g/mol | - | Q5VX84 | - |
Table 5. Pairwise alignment of PARP1 protein sequences (in distance from human) (HomoloGene:1222, NCBI)
| Gene Species | Gene Symbol | Identity (%) PROTEIN |
| vs. P.troglodytes | PARP1 | 99 |
| vs. M.mulatta | PARP1 | 98,2 |
| vs. C.lupus | PARP1 | 94,1 |
| vs. B.taurus | PARP1 | 90,4 |
| vs. M.musculus | Parp1 | 92,2 |
| vs. R.norvegicus | Parp1 | 91,6 |
| vs. G.gallus | PARP1 | 79,5 |
| vs. X.tropicalis | parp1 | 75,7 |
| vs. D.rerio | parp1 | 72,1 |
| vs. D.melanogaster | Parp | 43,8 |
| vs. A.gambiae | AgaP_AGAP003230 | 46,5 |
| vs. C.elegans | pme-1 | 41,1 |
| vs. A.thaliana | PARP1 | 42,3 |
| vs. O.sativa | Os07g0413700 | 42,7 |
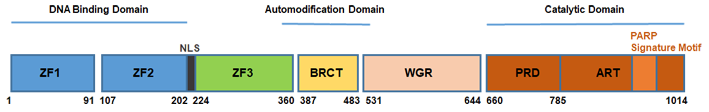
Description
PARP1 is known to be activated by mono-ADP-ribosylation, acetylation, increased cellular calcium concentration, or by binding to tyrosyl tRNA synthase. On the other hand, self-PARylation and sumoylation were shown to inhibit PARP1 activity. PARP1 can also be phosphorylated in a reversible manner and the phosphorylation can activate (e.g., AMP-activated protein kinase [AMPK]) or inhibit (e.g., protein kinase C) PARP-1 activity (Bai, 2015). In addition, physical interactions with other proteins, including histones, HPF1, HMGN1, XPA, NEIL1, OGG1, DDB2, TP53, and MAPK1 (ERK2) found to regulate PARP1 activity (Alemasova & Lavrik, 2019). It was also reported that dimerization of PARP1 enhances its enzymatic activity while further multimerization or dissociation to single PARP1 molecules leads to decreased enzymatic activity (Alemasova & Lavrik, 2019).
Expression
Localisation
PARP1 is one of several known cellular substrates of CASP3 and CASP7 (caspase 3 and caspase 7) and cleavage of PARP1 by these caspases is considered to be a hallmark of apoptosis. Upon cleavage, two specific fragments of PARP1 are generated: an 89 kDa fragment containing AMD and the catalytic domain of the enzyme and a 24 kDa containing DBD. The 89 kDa fragment has a greatly reduced DNA binding capacity and is liberated from the nucleus into the cytosol. On the other hand, the 24 kDa cleaved fragment with 2 zinc-finger motifs does not leave the nucleus where it binds to nicked DNA irreversibly and therefore acts as a trans-dominant inhibitor of active PARP1 (Chaitanya, Alexander, & Babu, 2010).
Function
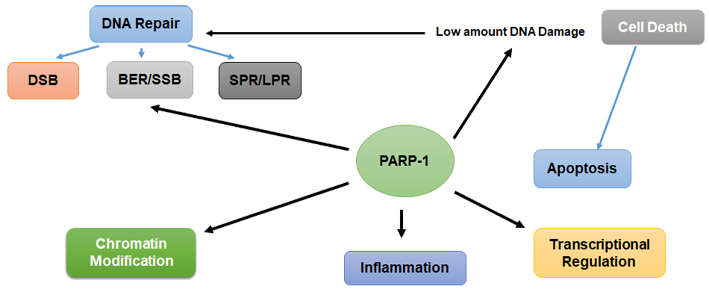
Although PARP1 has been long defined as a DNA-damage response protein, recent investigations highlight multiple functions of PARP1 including transcription, replication, aging, viral protection, cell cycle regulation, modification of chromosome structure, differentiation, inflammation, metabolic regulation, proteasomal degradation, and RNA processing (Bai, 2015; Rodrèguez-Vargas et al., 2019) (Figure 4).
PARP1 functions in DNA repair
Genotoxic stress results in various types of DNA lesions, including DNA single-strand breaks (SSBs) and double-strand breaks (DSBs). If not repaired, accumulated damage can disrupt genomic integrity. Fortunately, cells have evolved different DNA-damage repair responses that repair these DNA lesions to insure genomic stability (C. Liu, Vyas, Kassab, Singh, & Yu, 2017).
PARP1 recognizes both SSBs and DSBs and transfers the ADP-ribose moiety of NAD+ to the side chains of asparagine, aspartic acid, glutamic acid, arginine, lysine, serine and cysteine residues on its target proteins. Through their PAR-binding domains, these PAR chains form a platform and recruit DNA repair proteins. Therefore, PARP1 is an important DNA damage sensor for both SSBs and DSBs (Ray Chaudhuri & Nussenzweig, 2017).
PARP1 modulates chromatin structure and transcription
PARP1 functions in chromatin compaction, decondensation and it modulates epigenetic marks via PARylating histones and chromatin remodeling enzymes (Quénet, El Ramy, Schreiber, & Dantzer, 2009). Being as a component of enhancer/promoter-binding complexes, besides its effects on chromatin structure, PARP1 can bind to most of the RNA polymerase II transcribed genes and mediate around 3.5% of all transcribed RNAs covering a broad range of functions from inflammation to metabolism (Ke, Zhang, Lv, Zeng, & Ba, 2019; Kraus, 2008). PARP1 can also enhance the accessibility of promoters via histone and nucleosome replacements and can enhance transcription by replacing negative transcriptional cofactors with positive ones (Kraus & Hottiger, 2013; Muthurajan et al., 2014).
Recent findings described new roles of PARP1 in the regulation of RNA binding proteins, rRNA synthesis, ribosome biogenesis, and mRNA regulation (Ke et al., 2019; Ryu, Kim, & Kraus, 2015). Accordingly, PARP1 can regulate gene expression at the post-transcriptional level.
Cell death and PARP1
As mentioned before, PARP1 is known to be cleaved and inactivated by active caspases 3 and 7 and this cleavage is accepted as a hallmark of apoptosis (Castri et al., 2014; Desroches & Denault, 2019). The cleavage causes the formation of 24 kDa and 89 kDa fragments. Depending on the intensity and type of stimuli resulting in the cleavage, two main consequences have been reported: (1) reduced PARylation during DNA repair processes; (2) the modification of PARP1 transcriptional activity (Castri et al., 2014).
Recent studies indicate that PARP1 hyperactivation, ie. excessive PARylation by PARP1, can lead parthanatos, a form of necrotic cell death which PAR induces the nuclear translocation of apoptosis-inducing factor ( AIFM1) from mitochondria to initiate chromatinolysis and cell death independently of caspase activation. For a long time, it has been thought that the cell death caused by excessive activation of PARP occurs via the catalytic consumption of NAD+ followed by ATP reduction and bioenergetic collapse. However, Andrabi et al. showed that not the decreased NAD+, but PAR-dependent inhibition of hexokinase activity leads to defects in glycolysis and therefore causes the bioenergetic collapse. On the other hand, PARP1 activity is kept at much lower levels during normal unstressed cellular conditions (Andrabi et al., 2014; Dawicki-McKenna et al., 2015; Gupte, Liu, & Kraus, 2017).
Mutations
Note
Germinal
Somatic
Using genome-wide and high-density CRISPR-Cas9 tag-mutate-enrich mutagenesis screens, Pettitt et al. identified PARP1 mutant alleles that cause in vitro and in vivo PARP inhibitor resistance. The results reveal that point mutations in the ZnF domains were sufficient for the inhibitor resistance (Pettitt et al., 2018).
Polymorphisms: In addition to mutations, through modulation of PARP1 expression level and enzyme activity, PARP1 gene polymorphisms can affect the outcome and response to therapy of cancer. For example, PARP1 SNP rs1805407, found in perfect linkage disequilibrium with two PARP1 promoter SNPs (rs2077197 and rs6665208), was shown to be associated with higher PARP1 expression (Abecassis et al., 2019). In another study, expression quantitative trait locus (eQTL) analysis in melanocytic cell types revealed that presence of the 1q42.1 melanoma risk allele (rs3219090[G]) is correlated with higher PARP1 levels. Furthermore, a proteomic screen identified that RECQL helicase binds to the insertion allele of PARP1 (indel SNP rs144361550) in melanoma cells and primary human melanocytes (J. Choi et al., 2017). In another study, using a new data integrative approach applied on multi-modal -omics, and clinical data, Abecassis et al. demonstrated that response to chemotherapy is directly linked to the gene expression, four methylation variables and PARP1 SNP rs1805407 in a cohort of metastatic melanoma patients (Abecassis et al., 2019). According to the results of another genotyping study, Val762Ala, Asp81Asp, and Lys352Lys polymorphisms and the haplotype-ACAAC in PARP1 are associated with reduced risk of non-Hodgkin lymphoma in Korean males (Jin et al., 2010). In a case-control study conducted in the Hexi area of China, PARP1 2819G allele was shown to be associated with an increased risk of gastric cancer (He, Liu, Shan, Zhu, & Li, 2012).
In addition, PAR metabolism is also involved in malignancies. For instance, PARylation of proteins in peripheral blood leukocytes was shown to be reduced by more than 50% in head, neck, breast and cervical cancers (Lakadong, Kataki, & Sharan, 2010).
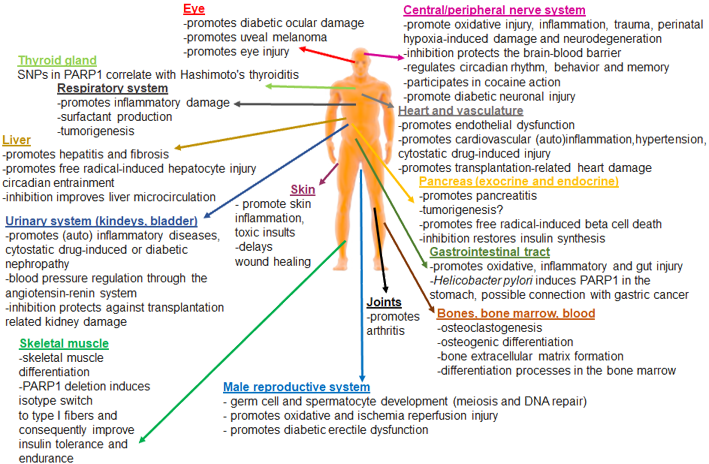
Implicated in
More recently, it has been shown that PARP1 can affect cardiac functions also via autophagy activation. Therefore, inhibition of PARP1 was suggested to be protective against cardiac ischemia injury by repressing autophagy (C. Wang, Xu, Zhang, Zhang, & Huang, 2018).
A growing number of evidence shows that mitochondrial function is strictly controlled by PARP1 which is responsible for about more than 90% of PARylation in the brain (Pieper et al., 2000). In addition to oxidative stress which is able to activate PARP1, recent studies claim that PARP1 is a critical component of a molecular interactions network responsible in the nervous system disorders related to mitochondrial function. It was suggested that deleterious consequences of PARP1 activation on mitochondrial function are caused by its interaction with SIRT1 (Sirtuin 1). In addition, the interaction of PARP1 with promoters of nuclear genes encoding for mitochondrial transcription factors and mtDNA repair proteins were identified (Czapski et al., 2018).
In addition to malignant tissues of BRCA-mutant, triple-negative, and receptor-positive breast carcinoma, PARP1 is overexpressed significantly in uterine carcinoma and ovarian carcinoma. As in breast carcinoma, ovarian cancer cells show high sensitivity to drugs designed for PARP1 inhibition (Iqbal et al., 2012; Thompson & Easton, 2003; L. Wang et al., 2017).
BCL6 is one of the therapeutic targets in lymphoma. As a transcription factor, BCL6 is expressed in germinal centre B cells and it is fundamental for the formation of germinal centres and the production of high-affinity antibodies. On the other hand, during terminal differentiation to plasma cells, BCL6 has to be transcriptionally downregulated. BCL6 is known to be highly expressed in B cell non-Hodgkins lymphoma and in a subset of cases of diffuse large cell lymphoma. PARP1 was shown to bind in a sequence-specific manner at the BCL6 locus and contributes to the regulation of BCL6 transcription (Ambrose, Papadopoulou, Beswick, & Wagner, 2007).
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 27890643 | 2017 | Coordination of DNA single strand break repair. | Abbotts R et al |
| 30824778 | 2019 | PARP1 rs1805407 Increases Sensitivity to PARP1 Inhibitors in Cancer Cells Suggesting an Improved Therapeutic Strategy. | Abecassis I et al |
| 31174925 | 2019 | PARP1: A potential biomarker for gastric cancer. | Afzal H et al |
| 30799503 | 2019 | Poly(ADP-ribosyl)ation by PARP1: reaction mechanism and regulatory proteins. | Alemasova EE et al |
| 24870775 | 2014 | Novel mutations of the PARP-1 gene associated with colorectal cancer in the Saudi population. | Alshammari AH et al |
| 19372272 | 2009 | Molecular mechanism of poly(ADP-ribosyl)ation by PARP1 and identification of lysine residues as ADP-ribose acceptor sites. | Altmeyer M et al |
| 17404575 | 2007 | Poly-(ADP-ribose) polymerase-1 (Parp-1) binds in a sequence-specific manner at the Bcl-6 locus and contributes to the regulation of Bcl-6 transcription. | Ambrose HE et al |
| 24987120 | 2014 | Poly(ADP-ribose) polymerase-dependent energy depletion occurs through inhibition of glycolysis. | Andrabi SA et al |
| 26091343 | 2015 | Biology of Poly(ADP-Ribose) Polymerases: The Factotums of Cell Maintenance. | Bai P et al |
| 10433620 | 1999 | Cancer risks in BRCA2 mutation carriers. | |
| 24333653 | 2014 | Poly(ADP-ribose) polymerase-1 and its cleavage products differentially modulate cellular protection through NF-kappaB-dependent signaling. | Castri P et al |
| 21176168 | 2010 | PARP-1 cleavage fragments: signatures of cell-death proteases in neurodegeneration. | Chaitanya GV et al |
| 26898760 | 2016 | PARP1 enhances lung adenocarcinoma metastasis by novel mechanisms independent of DNA repair. | Choi EB et al |
| 28759004 | 2017 | A common intronic variant of PARP1 confers melanoma risk and mediates melanocyte growth via regulation of MITF. | Choi J et al |
| 20942953 | 2010 | Evolutionary history of the poly(ADP-ribose) polymerase gene family in eukaryotes. | Citarelli M et al |
| 31133874 | 2019 | PARP Inhibitor PJ34 Protects Mitochondria and Induces DNA-Damage Mediated Apoptosis in Combination With Cisplatin or Temozolomide in B16F10 Melanoma Cells. | Cseh AM et al |
| 29128369 | 2018 | Inhibition of poly(ADP-ribose) polymerase-1 alters expression of mitochondria-related genes in PC12 cells: relevance to mitochondrial homeostasis in neurodegenerative disorders. | Czapski GA et al |
| 29632181 | 2018 | PARP-1 protects against colorectal tumor induction, but promotes inflammation-driven colorectal tumor progression. | Dörsam B et al |
| 28828398 | 2017 | Poly (ADP-Ribose) Polymerase-1 (PARP-1) Induction by Cocaine Is Post-Transcriptionally Regulated by miR-125b. | Dash S et al |
| 26626480 | 2015 | PARP-1 Activation Requires Local Unfolding of an Autoinhibitory Domain. | Dawicki-McKenna JM et al |
| 31586028 | 2019 | Caspase-7 uses RNA to enhance proteolysis of poly(ADP-ribose) polymerase 1 and other RNA-binding proteins. | Desroches A et al |
| 20338998 | 2010 | PARP1 deficiency exacerbates diet-induced obesity in mice. | Devalaraja-Narashimha K et al |
| 22937159 | 2012 | YY1-binding sites provide central switch functions in the PARP-1 gene expression network. | Doetsch M et al |
| 22053002 | 2012 | Poly(ADP-ribose)polymerase-1 (PARP1) controls adipogenic gene expression and adipocyte function. | Erener S et al |
| 9761745 | 1998 | Purification and biochemical characterization of a poly(ADP-ribose) polymerase-like enzyme from the thermophilic archaeon Sulfolobus solfataricus. | Faraone-Mennella MR et al |
| 21508395 | 2011 | Evidence for the efficacy of Iniparib, a PARP-1 inhibitor, in BRCA2-associated pancreatic cancer. | Fogelman DR et al |
| 22472897 | 2012 | PARP-1 protein expression in glioblastoma multiforme. | Galia A et al |
| 2109322 | 1990 | The second zinc-finger domain of poly(ADP-ribose) polymerase determines specificity for single-stranded breaks in DNA. | Gradwohl G et al |
| 16973716 | 2007 | Role of BRCA1 and BRCA2 mutations in pancreatic cancer. | Greer JB et al |
| 28202539 | 2017 | PARPs and ADP-ribosylation: recent advances linking molecular functions to biological outcomes. | Gupte R et al |
| 27028089 | 2016 | Poly(ADP-ribose) polymerase-1 silences retroviruses independently of viral DNA integration or heterochromatin formation. | Gutierrez DA et al |
| 11248084 | 2001 | Poly(ADP-ribose) polymerase-1 is required for efficient HIV-1 integration. | Ha HC et al |
| 22221038 | 2012 | PARP1 polymorphisms increase the risk of gastric cancer in a Chinese population. | He W et al |
| 23099806 | 2012 | The incidence of pancreatic cancer in BRCA1 and BRCA2 mutation carriers. | Iqbal J et al |
| 30672100 | 2019 | PARP inhibitors in ovarian cancer: Sensitivity prediction and resistance mechanisms. | Jiang X et al |
| 20196871 | 2010 | PARP-1 Val762Ala polymorphism is associated with reduced risk of non-Hodgkin lymphoma in Korean males. | Jin XM et al |
| 28279944 | 2017 | Adipose tissue NAD(+)-homeostasis, sirtuins and poly(ADP-ribose) polymerases -important players in mitochondrial metabolism and metabolic health. | Jokinen R et al |
| 30385548 | 2018 | Poly(ADP-ribose) drives pathologic α-synuclein neurodegeneration in Parkinson's disease. | Kam TI et al |
| 31055645 | 2019 | Novel insights into PARPs in gene expression: regulation of RNA metabolism. | Ke Y et al |
| 18450439 | 2008 | Transcriptional control by PARP-1: chromatin modulation, enhancer-binding, coregulation, and insulation. | Kraus WL et al |
| 23357755 | 2013 | PARP-1 and gene regulation: progress and puzzles. | Kraus WL et al |
| 31215481 | 2019 | MiR-7-5p-mediated downregulation of PARP1 impacts DNA homologous recombination repair and resistance to doxorubicin in small cell lung cancer. | Lai J et al |
| 29393407 | 2018 | PARP1-siRNA suppresses human prostate cancer cell growth and progression. | Lai Y et al |
| 21034502 | 2010 | ADP-ribose polymer--a novel and general biomarker of human cancers of head & neck, breast, and cervix. | Lakadong RO et al |
| 22582261 | 2012 | Structural basis for DNA damage-dependent poly(ADP-ribosyl)ation by human PARP-1. | Langelier MF et al |
| 28854736 | 2017 | The role of poly ADP-ribosylation in the first wave of DNA damage response. | Liu C et al |
| 27895737 | 2016 | High PARP-1 expression is associated with tumor invasion and poor prognosis in gastric cancer. | Liu Y et al |
| 28107648 | 2017 | PARP-1 Controls the Adipogenic Transcriptional Program by PARylating C/EBPβ and Modulating Its Transcriptional Activity. | Luo X et al |
| 25881554 | 2015 | PARP-1 involvement in neurodegeneration: A focus on Alzheimer's and Parkinson's diseases. | Martire S et al |
| 25136112 | 2014 | Automodification switches PARP-1 function from chromatin architectural protein to histone chaperone. | Muthurajan UM et al |
| 21779467 | 2010 | Upregulation of Poly (ADP-Ribose) Polymerase-1 (PARP1) in Triple-Negative Breast Cancer and Other Primary Human Tumor Types. | Ossovskaya V et al |
| 17919258 | 2007 | Role of poly(ADP-ribose) polymerase 1 (PARP-1) in cardiovascular diseases: the therapeutic potential of PARP inhibitors. | Pacher P et al |
| 30991935 | 2019 | ADP-ribosylation signalling and human disease. | Palazzo L et al |
| 31447348 | 2019 | LMO2 Confers Synthetic Lethality to PARP Inhibition in DLBCL. | Parvin S et al |
| 24865146 | 2014 | Distribution of protein poly(ADP-ribosyl)ation systems across all domains of life. | Perina D et al |
| 29748565 | 2018 | Genome-wide and high-density CRISPR-Cas9 screens identify point mutations in PARP1 causing PARP inhibitor resistance. | Pettitt SJ et al |
| 10677544 | 2000 | Poly(ADP-ribosyl)ation basally activated by DNA strand breaks reflects glutamate-nitric oxide neurotransmission. | Pieper AA et al |
| 18775502 | 2009 | The role of poly(ADP-ribosyl)ation in epigenetic events. | Quénet D et al |
| 19470756 | 2009 | SIRT1 promotes cell survival under stress by deacetylation-dependent deactivation of poly(ADP-ribose) polymerase 1. | Rajamohan SB et al |
| 28676700 | 2017 | The multifaceted roles of PARP1 in DNA repair and chromatin remodelling. | Ray Chaudhuri A et al |
| 31281570 | 2019 | PARP1 and Poly(ADP-ribosyl)ation Signaling during Autophagy in Response to Nutrient Deprivation. | Rodríguez-Vargas JM et al |
| 24916104 | 2014 | Transcriptional roles of PARP1 in cancer. | Schiewer MJ et al |
| 31217354 | 2019 | PARP-1 controls NK cell recruitment to the site of viral infection. | Shou Q et al |
| 2121735 | 1990 | Expression and site-directed mutagenesis of the catalytic domain of human poly(ADP-ribose)polymerase in Escherichia coli. Lysine 893 is critical for activity. | Simonin F et al |
| 24202328 | 2013 | PARP-1: Friend or Foe of DNA Damage and Repair in Tumorigenesis? | Swindall AF et al |
| 25378300 | 2014 | Opposing roles of mitochondrial and nuclear PARP1 in the regulation of mitochondrial and nuclear DNA integrity: implications for the regulation of mitochondrial function. | Szczesny B et al |
| 18452307 | 2008 | Domain C of human poly(ADP-ribose) polymerase-1 is important for enzyme activity and contains a novel zinc-ribbon motif. | Tao Z et al |
| 12237281 | 2002 | Cancer Incidence in BRCA1 mutation carriers. | Thompson D et al |
| 23416893 | 2013 | Poly(ADP-ribose) signaling in cell death. | Virág L et al |
| 23917125 | 2013 | A systematic analysis of the PARP protein family identifies new functions critical for cell physiology. | Vyas S et al |
| 30323296 | 2018 | PARP1 promote autophagy in cardiomyocytes via modulating FoxO3a transcription. | Wang C et al |
| 25412407 | 2015 | Inhibition of poly (ADP-ribose) polymerase and inducible nitric oxide synthase protects against ischemic myocardial damage by reduction of apoptosis. | Wang J et al |
| 28991194 | 2017 | PARP1 in Carcinomas and PARP1 Inhibitors as Antineoplastic Drugs. | Wang L et al |
| 31015836 | 2019 | Poly-ADP Ribosyl Polymerase 1 (PARP1) Regulates Influenza A Virus Polymerase. | Westera L et al |
| 28031413 | 2017 | Let-7 Status Is Crucial for PARP1 Expression in HER2-Overexpressing Breast Tumors. | Wielgos ME et al |
| 29949759 | 2018 | The PARP1-Siah1 Axis Controls HIV-1 Transcription and Expression of Siah1 Substrates. | Yu D et al |
| 21504625 | 2011 | Novel targeted therapeutics: inhibitors of MDM2, ALK and PARP. | Yuan Y et al |
Other Information
Locus ID:
NCBI: 142
MIM: 173870
HGNC: 270
Ensembl: ENSG00000143799
Variants:
dbSNP: 142
ClinVar: 142
TCGA: ENSG00000143799
COSMIC: PARP1
RNA/Proteins
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
PharmGKB
| Entity ID | Name | Type | Evidence | Association | PK | PD | PMIDs |
|---|---|---|---|---|---|---|---|
| PA166163418 | rucaparib | Chemical | LabelAnnotation | associated |
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 36448327 | 2024 | Functional genetic variation in 3'UTR of PARP1 indicates a decreased risk and a better severity of ischemic stroke. | 0 |
| 37714792 | 2024 | PARP1 roles in DNA repair and DNA replication: The basi(c)s of PARP inhibitor efficacy and resistance. | 3 |
| 38000246 | 2024 | Unraveling the allosteric inhibition mechanism of PARP-1 CAT and the D766/770A mutation effects via Gaussian accelerated molecular dynamics and Markov state model. | 1 |
| 38029688 | 2024 | Regulatory apoptotic fragment of PARP1 complements catalytic fragment for PAR and DNA-dependent activity but inhibits DNA-induced catalytic stimulation of PARP2. | 1 |
| 38043199 | 2024 | Methyltransferase Set7/9 controls PARP1 expression and regulates cisplatin response of breast cancer cells. | 1 |
| 38172525 | 2024 | Cancer-educated neutrophils promote lung cancer progression via PARP-1-ALOX5-mediated MMP-9 expression. | 0 |
| 38216672 | 2024 | TKT-PARP1 axis induces radioresistance by promoting DNA double-strand break repair in hepatocellular carcinoma. | 0 |
| 38297082 | 2024 | PARP1-stabilised FOXQ1 promotes ovarian cancer progression by activating the LAMB3/WNT/β-catenin signalling pathway. | 0 |
| 38320550 | 2024 | PARP1-DNA co-condensation drives DNA repair site assembly to prevent disjunction of broken DNA ends. | 4 |
| 38336156 | 2024 | A review of poly(ADP-ribose)polymerase-1 (PARP1) role and its inhibitors bearing pyrazole or indazole core for cancer therapy. | 2 |
| 38388665 | 2024 | Value of the NF-κB signalling pathway and the DNA repair gene PARP1 in predicting distant metastasis after breast cancer surgery. | 0 |
| 38391916 | 2024 | A Knockout of Poly(ADP-Ribose) Polymerase 1 in a Human Cell Line: An Influence on Base Excision Repair Reactions in Cellular Extracts. | 0 |
| 38521768 | 2024 | FANCJ promotes PARP1 activity during DNA replication that is essential in BRCA1 deficient cells. | 2 |
| 38593154 | 2024 | Blocking lipid synthesis induces DNA damage in prostate cancer and increases cell death caused by PARP inhibition. | 0 |
| 38657044 | 2024 | PARP1 UFMylation ensures the stability of stalled replication forks. | 0 |
Citation
Sinem Tunçer ; Kubra Kavak
PARP1 (poly(ADP-ribose) polymerase 1)
Atlas Genet Cytogenet Oncol Haematol. 2019-11-01
Online version: http://atlasgeneticsoncology.org/gene/586/parp1-(poly(adp-ribose)-polymerase-1)
