BCL6 (B-Cell Lymphoma 6)
2012-12-01 Jean-Loup Huret AffiliationGenetics, Dept Medical Information, University of Poitiers, CHU Poitiers Hospital, F-86021 Poitiers, France
Identity


DNA/RNA
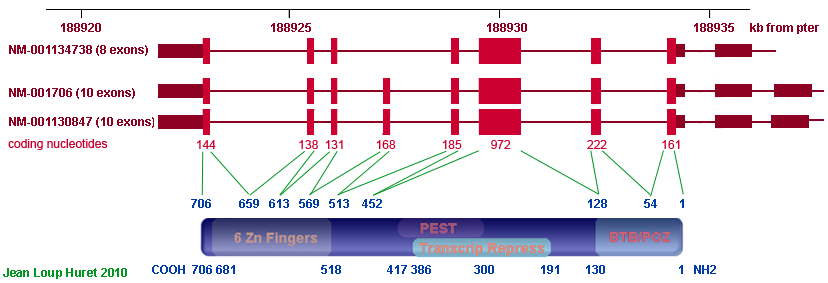
Description
Transcription
Proteins
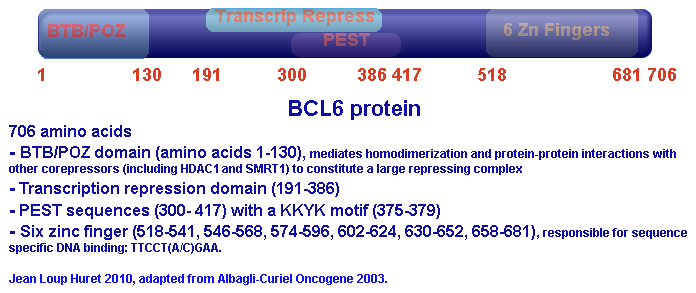
Description
Expression
Localisation
Function
The protein can bind to sequence specific DNA and repress its transcription in addition to recruiting other protein repressors. The DNA binding is mediated through the consensus sequence TTCCT(A/C)GAA (see below), while the protein-protein interactions are mediated through the BTB/POZ domain and it has been shown to interact with other zinc finger proteins and corepressors (including Histone Deacetylase 1 (HDAC1) and NCOR2 (Silencing Mediator of Retinoid and Thryoid Receptor 1 (SMRT1))). The carboxy terminus, on the other hand, is responsible for sequence specific DNA binding through its 6 zinc fingers.
Consensus DNA motifs recognized by BCL6
The Zn fingers of BCL6 binds to DNA and recognizes specific DNA motifs: the M00424 motif, a consensus sequence for the NKX-homeobox family of transcription factors, the M2 motif, which mimics an M00424 half site; and the M0 motif (see details in Basso et al., 2010).
BCL6 cellular function
- BCL6 cellular function in germinal centers of lymph nodes follicles
Mature naïve B cells migrate from the bone marrow to lymphoid nodes, meet the antigen, they are stimulated to proliferate and further differentiate into centroblasts in germinal centers (GC) of lymphoid follicles. There, cells are subjects to somatic hypermutation and class switch recombination, to further differenciate into memory B cells or plasma cells. This process also requires the presence of follicular helper T cells. BCL6 has a major role in this process. In particular, BCL6 prevents ATM and TP53 (Phan et al., 2004) to induce apoptosis in response to DNA rearrangements such as those necessary for somatic hypermutation and class switch recombination processes. BCL6 is therefore essential for normal B cell development (reviews in Albagli-Curiel, 2003; Jardin et al., 2007; Parekh et al., 2008; Basso and Dalla-Favera, 2010; Pellicano and Holyoake, 2011; Wagner et al., 2011; Basso and Dalla-Favera, 2012; Okada et al., 2012).
- T follicular helper cells
BCL6 is essential for the development of follicular helper T (Tfh) cells. Tfh cells are defined by the expression of the surface markers CXCR5 and PD-1 and synthesis of IL4 and IL21. BCL6 upregulation of B cells in the outer follicle sustains their interactions with helper T cells and allows them to enter the germinal centers (GC) region (Poholek et al., 2010; Kitano et al., 2011; Baumjohann et al., 2011). ICOS induces BCL6, and BCL6 then induces CXCR5. Tfh cells were lost in the absence of B cells, demonstrating a B cell requirement for maintenance of BCL6 and Tfh cell commitment (Choi et al., 2011). BCL6 and MAF cooperate to induce Tfh cell differentiation (Kroenke et al., 2012).
- Other tissues
BCL6 promotes survival of olfactory sensory neurons (Otaki etal., 2010).
BCL6 associate to NACC1 in brain regions (Korutla et al., 2009).
BCL6 forms a complex with BCOR on the promoters of Notch target genes in Xenopus embryos (Sakano et al., 2010).
BCL6-overexpression inhibits osteoclastogenesis, and the PRDM1 (Blimp-1)/BCL6 balance is essential in regulating the bone homeostasis by controlling osteoclastogenesis (Miyauchi et al., 2010).
BCL6 controls the expression of the CC-type chemokine genes and attenuates allergic airway inflammation in pulmonary epithelium (Seto et al. 2011).
BCL6 repression activity
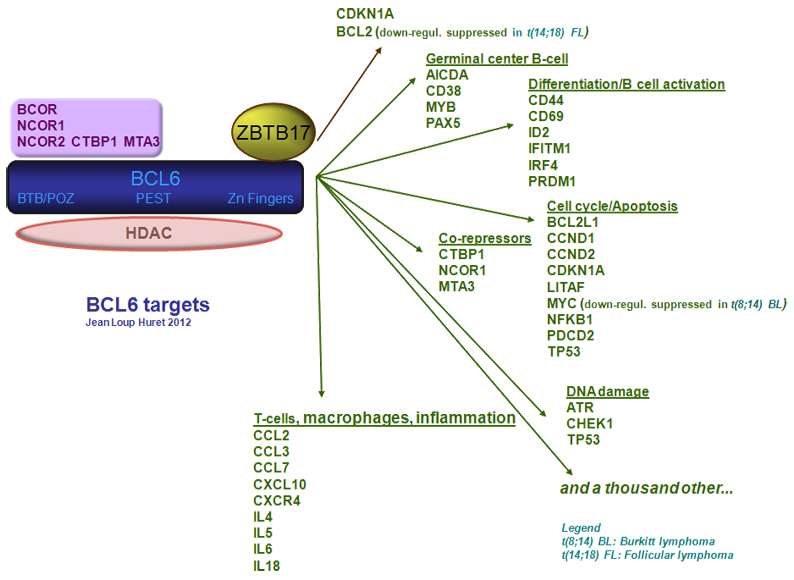
BCL6 binds to DNA as a homodimer and recruits co-repressor molecules, which, in turn, recruit class I and II histone deacetylases. BCL6 and histone deacetylases (HDACs) form stable complexes. BCL6 binds HDACs both directly through its C-terminal Zn fingers and through its N-terminal BTB/POZ domain and indirectly by recruiting several co-repressors such as BCOR, NCOR1, NCOR2 (which bind in a mutually exclusive way to the BTB/POZ domain), CTBP1 (which binds the BTB/POZ and the PEST domains), and MTA3 (which binds the PEST domain). Deacetylation of histones leads to transcriptional repression of BCL6 targets.
BCL6 interacts with ZBTB17 to repress CDKN1A and BCL2 (Phan et al., 2005). BCL6 has also been found to interact with other POK family members ((POZ and Krüppel)/ZBTB (zing finger and BTB) protein family) such as ZBTB7A (LRF) and ZBTB16 (PLZF).
BCL6 targets
An integrated biochemical and computational approach has recently identified thousands of BCL6 direct target genes, including B cell receptor (BCR; note: nothing to do with BCR, the partner of ABL1) and CD40 signaling genes (involved in signal transduction, MAPK activation, NF-AT activation, and NF-kB activation); T cell-mediated B-cell activation; apoptosis (BCL2, proteins involved in positive and negative regulation of the DISC complex and caspase activation, suggesting a role for BCL6 in balancing pro- and antiapoptotic programs in GC B cells); response to DNA damage (ATM, TP53, CDKN1A, ATR, CHEK1, TP53BP1...); interferon and cytokine signaling (interferon-type and interleukin receptors that lead to activation of JAK/STAT. STAT1, STAT3, and STAT5A were also found to be directly repressed by BCL6); Toll-like receptor signaling; TGFb receptor signaling; and WNT signaling (Basso et al., 2010).
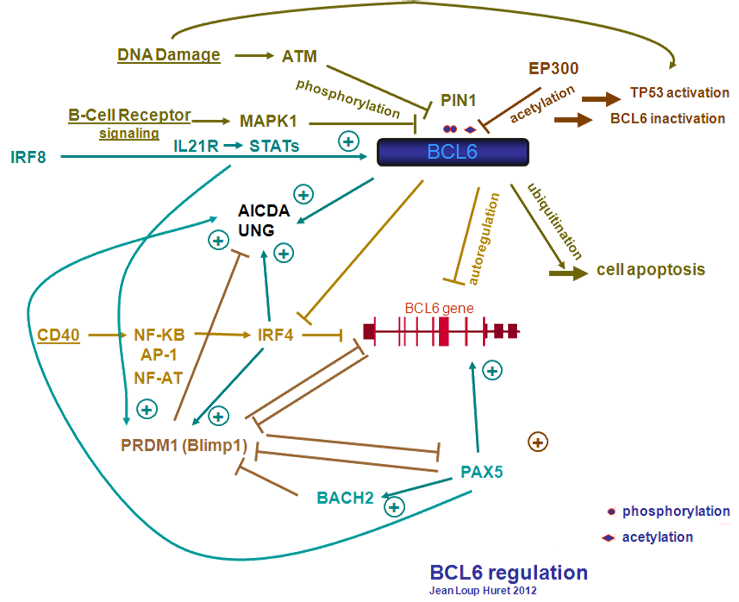
BCL6 regulation
- B cell receptor
B cell receptor (BCR) induces BCL6 phosphorylation by MAPK1 on Ser333 and Ser343, which targets BCL-6 for rapid degradation by the ubiquitin/proteasome pathway (Niu et al., 1998). FBXO11 promotes BCL6 ubiquitylation and degradation (Duan et al., 2012).
- DNA damage
After DNA damage, the kinase ATM promotes BCL6 phosphorylation; follows an interaction with PIN1 required for BCL6 degradation (Phan et al., 2007).
- EP300
EP300 binds and acetylates BCL6, which inactivates BCL6 (acetylation disrupts the ability of BCL6 to recruit histone deacetylases). The same two pathways (HDAC pathway and SIRT2 pathway) that regulate acetylation-mediated activation of TP53 also control acetylation-mediated inactivation of BCL6 (Bereshchenko et al., 2002). BCL6 represses the expression of EP300 and its cofactor BAT3 (Cerchietti et al., 2010). EP300 acetylates and activates PAX5 (He et al., 2011), which enhances BCL6.
- Autoregulation
BCL6 protein binds BCL6 gene promoter, and repress BCL6, mechanism of negative autoregulation (Pasqualucci et al., 2003). PATZ1 interacts with BCL6 and is required for its negative autoregulation (Pero et al., 2012).
- CD40 signaling
CD40 signaling activates various pathways, including NF-kB, NF-AT, and AP-1 (Francis et al., 1995). ILF2 (NF45) and ILF3 (NF90) form the nuclear factor of activated T-cells (NF-AT), a crucial transcription factor essential for productive T cell activation (Kao et al., 1994). CD40 signalling reduces BCL6 expression, through induction of IRF4 by the NF-kB pathway (Saito et al., 2007).
- PRDM1 (review in Crotty et al., 2010)
JUND/AP-1 and activated STATs drive high BCL6 expression in GC B cells (Arguni et al., 2006). STAT5 up-regulates BCL6 expression (Scheeren et al., 2005); in contrast, STAT3 up-regulates PRDM1 (Diehl et al., 2008) (the protein coded from PRDM1 is named Blimp-1). BCL6 directly promotes the expression of BACH2, which repress PRDM1 (Blimp-1). BCL6 also suppresses PRDM1 (Blimp-1) through repression of IRF4. IRF4 binds to and induces PRDM1. PRDM1 (Blimp-1) abrogates BCL6. PAX5 (protein name: BSAP) contributes to BCL6 expression (Nera et al., 2006; Tarlinton, 2011). PAX5 abrogates PRDM1 (Blimp-1), which abrogates PAX5. BCL6 and PAX5 promote the expression of AICDA and UNG (which have a major role in somatic hypermutation and class-switch recombination) (Alinikula et al., 2011). BCL6 positively regulates AICDA, IRF8, and MYB via repression of MIR155 (Basso et al., 2012). BCL6 mediates transcriptional repression by recruiting HDACs. MIR155 directly targets HDAC4 and indirectly attenuates BCL6 expression (Sandhu et al., 2012). PRDM1 (Blimp-1) represses BCL6, and also PAX5, MYC, and AICDA (review in Martins and Calame, 2008). IRF8 is expressed at the high levels in germinal center (GC) B cells. IRF8 binds BCL6 promoter and activates BCL6 (Lee et al., 2006).
It is also to be noted that IRF4 is involved in the t(6;14)(p25;q32) IRF4/IGH. PRDM1 (Blimp-1) is inactivated in about 50% of the activated B cell-like (ABC) subtype of diffuse large B cell lymphoma (ABC- DLBCL) (Mandelbaum et al., 2010).
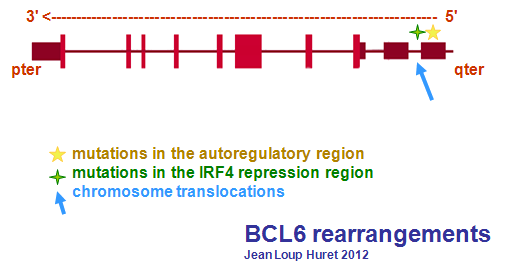
Mutations
Note
Mutations in the autoregulatory region of BCL6 leads to a constitutive BCL6 expression (Pasqualucci et al., 2003).
IRF4 response elements in the BCL6 gene mediate repression of BCL6. Mutations in the IRF4 repression region of BCL6 gene also leads to constitutive expression of BCL6 (Saito et al., 2007).
MYC, BCL2, and BCL6 rearrangements were detected in 6%, 15%, and 29% respectively of diffuse large B-cell lymphoma (DLBCL) patients. Double or triple rearrangements were detected in 3% of these patients. MYC rearrangement was associated with a significantly worse overall survival. BCL6 rearrangement also predicted a significantly shorter overall survival, especially for the non-GC phenotype (Akyurek et al., 2012).

Implicated in
- t(3;22)(q27;q11) the gene in 22q11 is IGL
In chronic myelogenous leukemia (CML), BCL6 acts as a critical effector downstream of FoxO in self-renewal signaling of CML-initiating cells. Therefore, pharmacological inhibition of BCL6 may represent a novel strategy to eradicate leukemia-initiating cells in CML (Hurtz et al., 2011).
Breakpoints
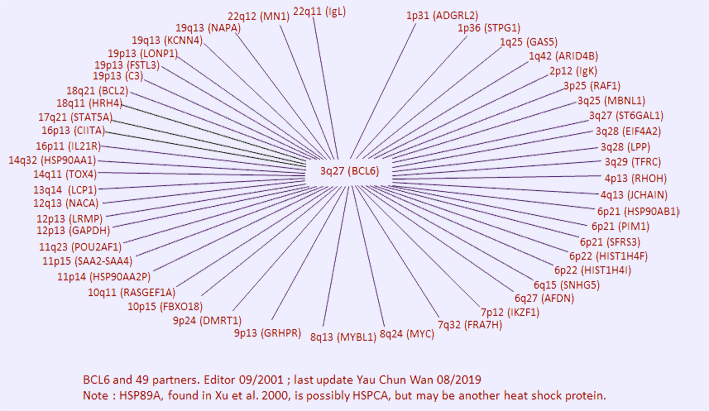
Note
Article Bibliography
| Pubmed ID | Last Year | Title | Authors |
|---|---|---|---|
| 22213394 | 2012 | Prognostic significance of MYC, BCL2, and BCL6 rearrangements in patients with diffuse large B-cell lymphoma treated with cyclophosphamide, doxorubicin, vincristine, and prednisone plus rituximab. | Akyurek N et al |
| 12555064 | 2003 | Ambivalent role of BCL6 in cell survival and transformation. | Albagli-Curiel O et al |
| 21674482 | 2011 | Alternate pathways for Bcl6-mediated regulation of B cell to plasma cell differentiation. | Alinikula J et al |
| 16702165 | 2006 | JunD/AP-1 and STAT3 are the major enhancer molecules for high Bcl6 expression in germinal center B cells. | Arguni E et al |
| 22500840 | 2012 | Roles of BCL6 in normal and transformed germinal center B cells. | Basso K et al |
| 19965633 | 2010 | Integrated biochemical and computational approach identifies BCL6 direct target genes controlling multiple pathways in normal germinal center B cells. | Basso K et al |
| 23166356 | 2012 | BCL6 positively regulates AID and germinal center gene expression via repression of miR-155. | Basso K et al |
| 21804014 | 2011 | Cutting Edge: Distinct waves of BCL6 expression during T follicular helper cell development. | Baumjohann D et al |
| 12402037 | 2002 | Acetylation inactivates the transcriptional repressor BCL6. | Bereshchenko OR et al |
| 21041953 | 2010 | BCL6 repression of EP300 in human diffuse large B cell lymphoma cells provides a basis for rational combinatorial therapy. | Cerchietti LC et al |
| 19701497 | 2009 | Bcl6 is expressed in neuroblastoma: tumor cell type-specific expression predicts outcome. | Chamdin A et al |
| 16772602 | 2006 | High BCL6 expression predicts better prognosis, independent of BCL6 translocation status, translocation partner, or BCL6-deregulating mutations, in gastric lymphoma. | Chen YW et al |
| 17542409 | 2007 | BCL6 gene mutations in transitional cell carcinomas. | Cho HY et al |
| 21636296 | 2011 | ICOS receptor instructs T follicular helper cell versus effector cell differentiation via induction of the transcriptional repressor Bcl6. | Choi YS et al |
| 20084069 | 2010 | Effectors and memories: Bcl-6 and Blimp-1 in T and B lymphocyte differentiation. | Crotty S et al |
| 18354204 | 2008 | STAT3-mediated up-regulation of BLIMP1 Is coordinated with BCL6 down-regulation to control human plasma cell differentiation. | Diehl SA et al |
| 22113614 | 2012 | FBXO11 targets BCL6 for degradation and is inactivated in diffuse large B-cell lymphomas. | Duan S et al |
| 21593872 | 2011 | BCL6 enables Ph+ acute lymphoblastic leukaemia cells to survive BCR-ABL1 kinase inhibition. | Duy C et al |
| 7537532 | 1995 | Induction of the transcription factors NF-kappa B, AP-1 and NF-AT during B cell stimulation through the CD40 receptor. | Francis DA et al |
| 21357426 | 2011 | Histone acetyltransferase p300 acetylates Pax5 and strongly enhances Pax5-mediated transcriptional activity. | He T et al |
| 20228272 | 2010 | A proto-oncogene BCL6 is up-regulated in the bone marrow microenvironment in multiple myeloma cells. | Hideshima T et al |
| 19337254 | 2009 | BCL6 degradation caused by the interaction with the C-terminus of pro-HB-EGF induces cyclin D2 expression in gastric cancers. | Hirata Y et al |
| 21911423 | 2011 | BCL6-mediated repression of p53 is critical for leukemia stem cell survival in chronic myeloid leukemia. | Hurtz C et al |
| 7519613 | 1994 | Cloning and expression of cyclosporin A- and FK506-sensitive nuclear factor of activated T-cells: NF45 and NF90. | Kao PN et al |
| 16867873 | 2006 | Intrachromosomal rearrangement of chromosome 3q27: an under recognized mechanism of BCL6 translocation in B-cell non-Hodgkin lymphoma. | Keller CE et al |
| 8220427 | 1993 | LAZ3, a novel zinc-finger encoding gene, is disrupted by recurring chromosome 3q27 translocations in human lymphomas. | Kerckaert JP et al |
| 21636294 | 2011 | Bcl6 protein expression shapes pre-germinal center B cell dynamics and follicular helper T cell heterogeneity. | Kitano M et al |
| 19121354 | 2009 | NAC1, a POZ/BTB protein that functions as a corepressor. | Korutla L et al |
| 22427637 | 2012 | Bcl6 and Maf cooperate to instruct human follicular helper CD4 T cell differentiation. | Kroenke MA et al |
| 16380510 | 2006 | Regulation of the germinal center gene program by interferon (IFN) regulatory factor 8/IFN consensus sequence-binding protein. | Lee CH et al |
| 12944904 | 2003 | BCL-6 is expressed in breast cancer and prevents mammary epithelial differentiation. | Logarajah S et al |
| 21156281 | 2010 | BLIMP1 is a tumor suppressor gene frequently disrupted in activated B cell-like diffuse large B cell lymphoma. | Mandelbaum J et al |
| 18370921 | 2008 | Regulation and functions of Blimp-1 in T and B lymphocytes. | Martins G et al |
| 8274740 | 1994 | Gene involved in the 3q27 translocation associated with B-cell lymphoma, BCL5, encodes a Krüppel-like zinc-finger protein. | Miki T et al |
| 20368579 | 2010 | The Blimp1-Bcl6 axis is critical to regulate osteoclast differentiation and bone homeostasis. | Miyauchi Y et al |
| 16546097 | 2006 | Loss of Pax5 promotes plasma cell differentiation. | Nera KP et al |
| 17197022 | 2007 | Prognostic impact of chromosomal alteration of 3q27 on nodal B-cell lymphoma: correlation with histology, immunophenotype, karyotype, and clinical outcome in 329 consecutive patients. | Niitsu N et al |
| 9649500 | 1998 | Antigen receptor signaling induces MAP kinase-mediated phosphorylation and degradation of the BCL-6 transcription factor. | Niu H et al |
| 17142954 | 2006 | Pathogenetic and clinical implications of non-immunoglobulin ; BCL6 translocations in B-cell non-Hodgkin's lymphoma. | Ohno H et al |
| 22500836 | 2012 | Differentiation of germinal center B cells and follicular helper T cells as viewed by tracking Bcl6 expression dynamics. | Okada T et al |
| 20151461 | 2010 | The proto-oncogene BCL6 promotes survival of olfactory sensory neurons. | Otaki JM et al |
| 18452090 | 2008 | Therapeutic targeting of the BCL6 oncogene for diffuse large B-cell lymphomas. | Parekh S et al |
| 12515714 | 2003 | Mutations of the BCL6 proto-oncogene disrupt its negative autoregulation in diffuse large B-cell lymphoma. | Pasqualucci L et al |
| 22025499 | 2011 | Assembling defenses against therapy-resistant leukemic stem cells: Bcl6 joins the ranks. | Pellicano F et al |
| 22493480 | 2012 | POZ-, AT-hook-, and zinc finger-containing protein (PATZ) interacts with human oncogene B cell lymphoma 6 (BCL6) and is required for its negative autoregulation. | Pero R et al |
| 15577913 | 2004 | The BCL6 proto-oncogene suppresses p53 expression in germinal-centre B cells. | Phan RT et al |
| 17828269 | 2007 | Genotoxic stress regulates expression of the proto-oncogene Bcl6 in germinal center B cells. | Phan RT et al |
| 20519643 | 2010 | In vivo regulation of Bcl6 and T follicular helper cell development. | Poholek AC et al |
| 17785208 | 2007 | A signaling pathway mediating downregulation of BCL6 in germinal center B cells is blocked by BCL6 gene alterations in B cell lymphoma. | Saito M et al |
| 20230751 | 2010 | BCL6 canalizes Notch-dependent transcription, excluding Mastermind-like1 from selected target genes during left-right patterning. | Sakano D et al |
| 15711548 | 2005 | STAT5 regulates the self-renewal capacity and differentiation of human memory B cells and controls Bcl-6 expression. | Scheeren FA et al |
| 21801248 | 2011 | Bcl6 in pulmonary epithelium coordinately controls the expression of the CC-type chemokine genes and attenuates allergic airway inflammation. | Seto T et al |
| 17175383 | 2007 | A novel t(2;3)(p11;q27) in a case of follicular lymphoma. | Tapinassi C et al |
| 21792874 | 2011 | Bcl6: where too much complexity is barely enough. | Tarlinton DM et al |
| 20124477 | 2010 | Prolactin inhibits BCL6 expression in breast cancer through a Stat5a-dependent mechanism. | Tran TH et al |
| 21083654 | 2011 | The role of BCL6 in lymphomas and routes to therapy. | Wagner SD et al |
| 21725042 | 2011 | BCL-6 expression in mesenchymal tumours: an immunohistochemical and fluorescence in situ hybridisation study. | Walters MP et al |
| 17175379 | 2007 | A novel t(3;8)(q27;q24.1) simultaneously involving both the BCL6 and MYC genes in a diffuse large B-cell lymphoma. | Wang HY et al |
| 8235596 | 1993 | Alterations of a zinc finger-encoding gene, BCL-6, in diffuse large-cell lymphoma. | Ye BH et al |
Other Information
Locus ID:
NCBI: 604
MIM: 109565
HGNC: 1001
Ensembl: ENSG00000113916
Variants:
dbSNP: 604
ClinVar: 604
TCGA: ENSG00000113916
COSMIC: BCL6
RNA/Proteins
Expression (GTEx)
Pathways
Protein levels (Protein atlas)
References
| Pubmed ID | Year | Title | Citations |
|---|---|---|---|
| 37897298 | 2024 | Alternative genetic alterations of MYC, BCL2, and/or BCL6 in high-grade B-cell lymphoma (HGBL) and diffuse large B-cell lymphoma (DLBCL): Can we identify different prognostic subgroups? | 0 |
| 38196006 | 2024 | Delineating the mechanism of fragility at BCL6 breakpoint region associated with translocations in diffuse large B cell lymphoma. | 0 |
| 37897298 | 2024 | Alternative genetic alterations of MYC, BCL2, and/or BCL6 in high-grade B-cell lymphoma (HGBL) and diffuse large B-cell lymphoma (DLBCL): Can we identify different prognostic subgroups? | 0 |
| 38196006 | 2024 | Delineating the mechanism of fragility at BCL6 breakpoint region associated with translocations in diffuse large B cell lymphoma. | 0 |
| 37085097 | 2023 | B-cell lymphoma 6 expression significantly differs by the uterine preparation method used for frozen embryo transfer. | 1 |
| 37140301 | 2023 | Clinical course and features of persistent polyclonal B-cell lymphocytosis with BCL-6 amplification during pregnancy. | 0 |
| 37406588 | 2023 | Identification of mutation gene prognostic biomarker in multiple myeloma through gene panel exome sequencing and transcriptome analysis in Chinese population. | 1 |
| 37950867 | 2023 | Bcl6 is a subset-defining transcription factor of lymphoid tissue inducer-like ILC3. | 1 |
| 37085097 | 2023 | B-cell lymphoma 6 expression significantly differs by the uterine preparation method used for frozen embryo transfer. | 1 |
| 37140301 | 2023 | Clinical course and features of persistent polyclonal B-cell lymphocytosis with BCL-6 amplification during pregnancy. | 0 |
| 37406588 | 2023 | Identification of mutation gene prognostic biomarker in multiple myeloma through gene panel exome sequencing and transcriptome analysis in Chinese population. | 1 |
| 37950867 | 2023 | Bcl6 is a subset-defining transcription factor of lymphoid tissue inducer-like ILC3. | 1 |
| 34654055 | 2022 | Diffuse large B-cell lymphomas in adults with aberrant coexpression of CD10, BCL6, and MUM1 are enriched in IRF4 rearrangements. | 13 |
| 34809978 | 2022 | B-cell lymphoma 6 expression is not associated with live birth in a normal responder in vitro fertilization population. | 2 |
| 35039330 | 2022 | Defective STAT5 Activation and Aberrant Expression of BCL6 in Naive CD4 T Cells Enhances Follicular Th Cell-like Differentiation in Patients with Granulomatosis with Polyangiitis. | 6 |
Citation
Jean-Loup Huret
BCL6 (B-Cell Lymphoma 6)
Atlas Genet Cytogenet Oncol Haematol. 2012-12-01
Online version: http://atlasgeneticsoncology.org/gene/20/bcl6-(b-cell-lymphoma-6)
Historical Card
2007-02-01 BCL6 (B-Cell Lymphoma 6) by Stevan Knezevich Affiliation
1998-09-01 BCL6 (B-Cell Lymphoma 6) by Jean-Pierre Kerckaert Affiliation
